BREAKING NEWS! Cardiff University Researchers Uncover How SARS-CoV-2 Evades CD4+ T Cell Immunity Via HLA Class II-Presented Epitopes
COVID-19 News - SARS-CoV-2 Evades CD4+ T Cell Immunity - HLA Class II-Presented Epitopes Jul 20, 2023 2 years, 10 months, 2 days, 4 hours, 28 minutes ago
Specifically, the Omicron variant of SARS-CoV-2 was found to escape pre-existing immune memory to two epitopes through two distinct mechanisms.
COVID-19 News: The ongoing battle against the COVID-19 pandemic relies not only on vaccines and antibodies but also on the critical role played by CD4+ T cells in providing long-lasting immunity. These T cells recognize specific peptide epitopes of the SARS-CoV-2 virus, contributing to immune memory and protection against severe disease.
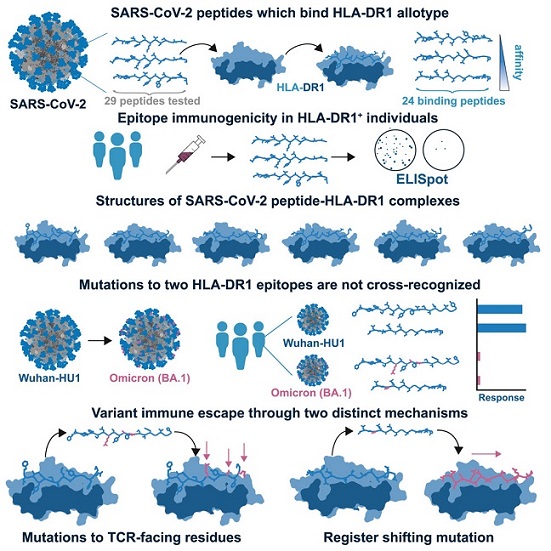 Graphical Abstract
Graphical Abstract
However, a recent study conducted by researchers at Cardiff University in the United Kingdom has revealed a startling discovery: the immunogenicity of these SARS-CoV-2 epitopes does not correlate with their binding affinity to the human leukocyte antigen class II (HLA-II) molecules. Furthermore, the study sheds light on how the virus employs sophisticated mechanisms to escape pre-existing CD4+ T cell immunity. These findings have significant implications for understanding immune responses to SARS-CoV-2 and developing effective strategies to combat the virus.
Exploring CD4+ T Cell Epitopes
CD4+ T cells play a crucial role in recognizing and eliminating viral infections. In the case of SARS-CoV-2, these T cells recognize a diverse range of peptide epitopes derived from various viral proteins. Previous studies and
COVID-19 News reports have shown that CD4+ T cell immunity contributes to protection against reinfection and severe COVID-19. However, the precise characterization of specific epitopes presented on the HLA-II molecules to CD4+ T cells has been limited.
The Cardiff University study aimed to bridge this knowledge gap by focusing on HLA-DR1, a common HLA-II variant. The study team assessed the binding affinity of 29 candidate peptides derived from SARS-CoV-2 to HLA-DR1 and characterized their immunogenicity in vitro.
Surprisingly, they discovered that the immunogenicity of these peptides did not correlate with their binding affinity, challenging previous assumptions about the importance of peptide-HLA-II interactions.
Structural Insights into Immune Escape
To understand the molecular basis of immune escape, the study team studied X-ray crystallographic structures of six epitopes bound to HLA-DR1. This structural analysis revealed how viral variant mutations impact epitope presentation.
Specifically, the Omicron variant of SARS-CoV-2 was found to escape pre-existing immune memory to two epitopes through two distinct mechanisms. One mechanism involved mutations to epitope positions directly recognized by T cell receptors (TCRs). The other mechanism, more s
ubtle yet equally effective, involved a single amino acid substitution that caused a register shift within the HLA binding groove, fundamentally altering the peptide-HLA structure.
Implications for Vaccine Design and Immune Protection
These findings have important implications for vaccine design and our understanding of SARS-CoV-2 immune responses. The study highlights the need to consider the unique characteristics of peptide-HLA-II interactions when developing vaccines and designing therapies to induce robust CD4+ T cell responses. By targeting more conserved regions of the viral proteome, which exhibit higher homology across different variants, it may be possible to induce stronger and broader immune responses that provide cross-protection against heterologous viruses.
The study also emphasizes the complexity of viral evolution and its impact on CD4+ T cell recognition. Viral variants can employ sophisticated strategies to evade immune responses, making it crucial to monitor and adapt vaccination strategies to counter these emerging threats. The researchers suggest that further investigation into T cell receptor clonotypes and their binding modes will provide deeper insights into the mechanisms underlying immune escape.
Conclusion
The Cardiff University study has provided significant insights into the mechanisms employed by SARS-CoV-2 to escape pre-existing CD4+ T cell immunity. The findings challenge conventional assumptions about the correlation between peptide-HLA-II binding affinity and immunogenicity. The study underscores the importance of considering the structural characteristics of peptide-HLA-II interactions when designing vaccines and therapies. By understanding how viral variants evade CD4+ T cell recognition, researchers can develop more effective strategies to combat COVID-19 and potentially other viral infections. Continued research in this field will pave the way for improved vaccines and treatments that harness the full potential of CD4+ T cell immunity.
The study findings were published in the peer reviewed journal: Cell Reports.
https://www.cell.com/cell-reports/fulltext/S2211-1247(23)00838-0
For the latest
COVID-19 News, keep on logging to Thailand Medical News.
