COVID-19 News: Researchers Unravel The Enigmatic Evolutionary Journey of SARS-CoV-2 Nonstructural Protein 6 (NSP6)
COVID-19 News – SARS-CoV-2 Nonstructural Protein 6 - NSP6 Jul 24, 2023 2 years, 9 months, 1 week, 4 days, 21 hours, 48 minutes ago
COVID-19 News: Since its first appearance in late 2019, the SARS-CoV-2 virus has embarked on a relentless evolutionary journey, presenting humanity with an ever-changing array of variants and challenges. As the virus adapts and mutates, it raises concerns about vaccine efficacy, therapeutic strategies, and the potential for new waves of infections. Among the enigmatic components influencing the behavior of SARS-CoV-2 variants, the nonstructural protein 6 (nsp6) has recently emerged as a key player in the virus's pathogenesis. New research has shed light on the fascinating evolution of this transmembrane protein, unveiling hidden secrets that could shape our battle against the pandemic.
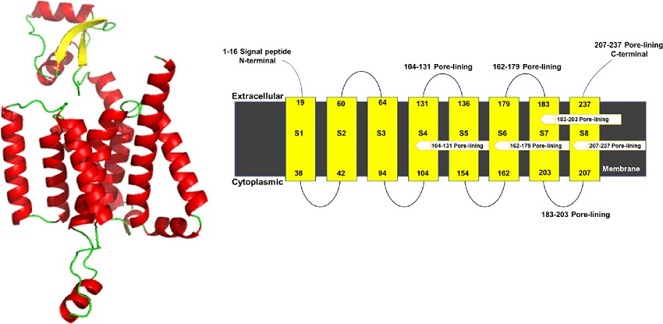 Three-dimensional model structure of SARS-CoV-2 NSP6 generated by AlphaFold. The transmembrane helices predicted through THHMM server (Credit: Kartikay Prasad)
Three-dimensional model structure of SARS-CoV-2 NSP6 generated by AlphaFold. The transmembrane helices predicted through THHMM server (Credit: Kartikay Prasad)
Numerous past
COVID-19 News reports have also covered on possible mechanisms and influences of the NSP6 protein.
https://www.thailandmedical.news/news/covid-19-news-university-of-texas-scientists-uncover-multiple-roles-of-nsp6-in-sars-cov-2-pathogenesis
https://www.thailandmedical.news/news/breaking-mutations-on-omicron-non-structural-protein-6-nsp6-alters-viral-disruption-modes-of-autophagy-in-host,-leading-to-possibly-more-serious-long-
https://www.thailandmedical.news/news/the-interplay-between-sars-cov-2-and-the-endoplasmic-reticulum-and-the-resulting-covid-19-symptoms-and-long-covid-manifestations
One alarming past study finding was that NSP6 proteins could actually cause morphological changes and functional defects to the human heart!
https://www.thailandmedical.news/news/breaking-university-of-maryland-study-shockingly-finds-that-sars-cov-2-nsp6-proteins-can-induce-morphological-and-functional-defects-in-the-heart
https://www.thailandmedical.news/news/breaking-indiana-university-finds-that-transcriptome-of-human-pluripotent-stem-cell-derived-cardiomyocytes-compromised-by-sars-cov-2-nsp6,-nsp8-and-m-
&
nbsp;
Unraveling the Role of NSP6
Scientists had previously focused on mutations in the spike glycoprotein, which allows the virus to escape neutralizing antibodies and adopt alternative pathways for infection. However, the role of nsp6 mutations remained less clear until a groundbreaking discovery came to light. Recent studies revealed that nsp6 is essential for modifying the endoplasmic reticulum and generating double-membrane vesicles (DMVs), the site of viral RNA replication. These DMVs play a crucial role in the virus's replication and spread within the host, making nsp6 an intriguing target for investigation.
A Genetic Expedition - Tracing the Footprints of NSP6 Mutations
To understand the evolution of nsp6, a group of dedicated researchers from Loyola University Chicago-USA and Boston University Chobanian and Avedisian School of Medicine-USA embarked on a vast genetic expedition. They analyzed a staggering dataset of 91,596 high-confidence human SARS-CoV-2 whole-genome sequences across 19 variants and lineages, spanning the entire timeline of the pandemic. Their findings provided a window into the changing landscape of nsp6 mutations.
Intriguingly, the researchers discovered that early variants of concern, including Alpha, Beta, and Gamma, carried a triple amino acid deletion (106–108, termed ΔSGF) in nsp6. In stark contrast, the more recent Delta, Epsilon, and Mu lineages retained the ancestral nsp6 sequence, presenting an evolutionary divergence in this essential protein.
The Rise of Omicron - The Mystery Deepens
As the SARS-CoV-2 virus continued its march across the globe, the emergence of the Omicron variant grabbed global attention. The researchers unraveled a fascinating transition in nsp6 mutations within the Omicron lineage. While Omicron BA.1 presented an amino acid 105–107 ΔLSG deletion in nsp6, the subsequent lineages, including BA.2 and beyond, demonstrated increased dominance of the ΔSGF deletion.
This shift in nsp6 mutations within Omicron variants piqued the curiosity of scientists. The prevalence of the ΔSGF deletion in multiple, independent variants suggested that SARS-CoV-2 nsp6 ΔSGF deletions might have been independently selected during the pandemic's course for viral adaptation in humans.
However, a puzzling observation emerged when comparing nsp6 sequences from SARS-CoV-2-related coronaviruses in bats and pangolins, which showed no sign of the ΔSGF/ΔLSG deletions. This finding suggested that the deletion in nsp6 might be a specific adaptation to human hosts.
Unveiling the Structure - The Hidden Architecture of NSP6
Unlocking the structure and function of nsp6 became the next frontier for researchers. Computational analyses using various tools provided intriguing insights into the predicted topology of SARS-CoV-2 nsp6. It emerged as a multipass transmembrane protein with a conserved C-terminal domain, hinting at its essential role in the virus's lifecycle. The location of the ΔSGF/ΔLSG deletion in the predicted structure raised questions about how these mutations influenced the protein's overall function and viral replication.
Implications and Future Directions
As the COVID-19 pandemic persists, the knowledge gained from studying nsp6 is proving invaluable. By understanding the factors contributing to the evolution and pathogenesis of SARS-CoV-2, researchers can better prepare for potential future variants and tailor effective interventions.
The quest to unlock the secrets of nsp6 continues, with researchers focusing on identifying host factors interacting with the protein during membrane zippering and DMV formation. Such discoveries could provide critical clues for developing targeted therapies aimed at limiting viral replication and mitigating the impact of emerging coronaviruses.
Conclusion
The SARS-CoV-2 emergence, marked by twists and turns, has been a rollercoaster of challenges and discoveries. The nonstructural protein 6, once an enigma, has emerged as a pivotal player in the virus's evolution and pathogenesis. More detailed research is required to find ways to inhibit its mechanisms using safe therapeutics.
The study findings were published in the peer reviewed journal: mBIO (ASM)
https://journals.asm.org/doi/10.1128/mbio.00688-23
For the latest
COVID-19 News, keep on logging to Thailand Medical News
