COVID-19 News: SARS-CoV-2 NSP3 Hijacks Fragile X Mental Retardation Proteins For Efficient Infection While Modulating Stress Granule Function!
Nikhil Prasad Fact checked by:Thailand Medical News Team Sep 06, 2023 2 years, 9 months, 2 days, 4 hours, 6 minutes ago
COVID-19 News: The COVID-19 pandemic has brought the world to its knees, prompting extensive research into the SARS-CoV-2 virus and its intricate interactions with host cells. A recent study conducted collaboratively by the University of Copenhagen-Denmark, the University of Texas Medical Branch-USA, and Umeå University-Sweden has shed new light on how this novel coronavirus manipulates host factors for efficient infection while modulating stress granule function.
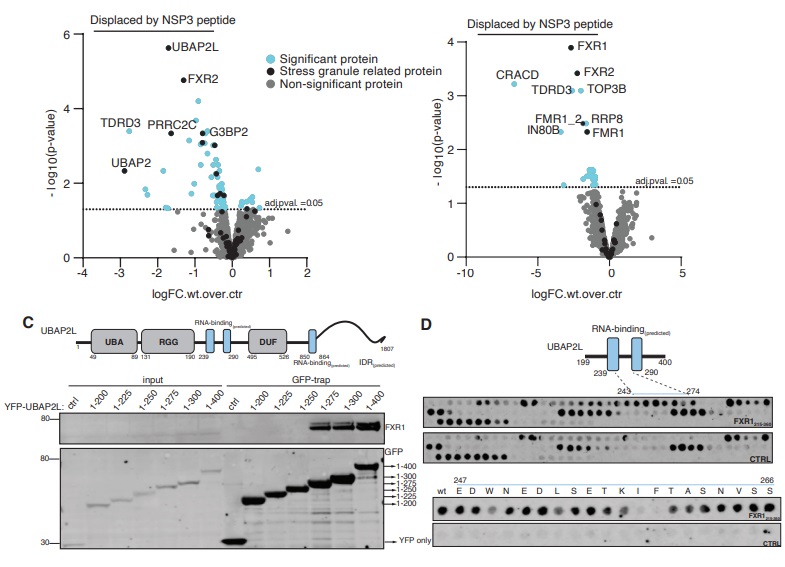 NSP3 disrupts the UBAP2L-FMRP complex.
NSP3 disrupts the UBAP2L-FMRP complex.
A) FXR1 was affinity purified and incubated with either WT NSP3 or mutant NSP3 peptide and interactomes determined by MS to determined proteins specifically displaced by WT NSP3. Data from 4 technical repeats. B) As A) but using UBAP2L as a bait. Data from 4 technical repeats. C) Schematic of UBAP2L and truncation analysis to identify FXR1 binding site (n=1). D) Peptide array of UBAP2L 199-400 to identify FXR1 binding region and lower part single Ala scan through UBAP2L 247-266 to identify critical residues (n=1).
The study reveals a previously unknown interaction between SARS-CoV-2's NSP3 protein and Fragile X Mental Retardation Proteins (FMRPs), a discovery that could have far-reaching implications for our understanding of both viral infection and fragile X syndrome.
The Complexity of Viral-Host Interactions
Viruses, despite their simplicity in terms of genetic material, rely heavily on their ability to interact with cellular host factors to replicate within host cells. Moreover, they possess mechanisms to dampen the host's innate immune responses, ensuring their successful proliferation as covered in previous studies and also
COVID-19 News reports. One of the intriguing targets of viruses is the manipulation of stress granules, which serve as vital hubs in the host's antiviral defense mechanisms.
Stress granules, intricate protein-RNA assemblies devoid of membranes, form in the cytoplasm as a response to various stress signals, including viral infections. Stress granules have long been proposed to have a function to protect RNAs from harmful conditions, thus their appearance under stress. The accumulation of RNAs into dense globules could keep them from reacting with harmful chemicals and safeguard the information coded in their RNA sequence.
These granules, while primarily associated with sequestering RNA, are also composed of a myriad of RNA-binding proteins, such as G3BP1/2 and UBAP2L, which play pivotal roles in nucleating and coordinating stress granule formation.
Remarkably, more than 250 host proteins have been implicated in the complex process of stress granule formation, underscoring its multifaceted nature. Recognizing the significance of stress granules in antiviral defense, viruses have developed strategies to disrupt their formation and even co-opt these factors to enhance their replication.
Several viral proteins, including the nucleocapsid protein (N) and accessory ORFs of coronaviruses, have been identified as disrupting stress granule f
ormation. Recently, it was revealed that SARS-CoV-2's N protein contains a specific motif that binds to G3BP1/2, effectively disrupting stress granules. This represents one of the virus's tactics to counter cellular antiviral mechanisms. Furthermore, both old and new world alphaviruses employ similar strategies by binding to G3BP1/2 through specific motifs in their NSP3 proteins, thereby facilitating viral replication complex assembly and disrupting stress granules. Intriguingly, Eastern equine encephalitis virus (EEEV) targets both G3BP1/2 and FMRPs, RNA-binding proteins, through distinct sequences in its hypervariable domain. FMRPs, known for their role in stress granule assembly, are also implicated in fragile X syndrome, a common form of inherited mental retardation. This connection underscores the importance of stress granules in host functions and hints at potential insights into the molecular basis of fragile X syndrome. In this study, the study team explored the novel interaction between SARS-CoV-2 NSP3 and FMRPs, shedding light on its role during viral infection.
Revelation of the SARS-CoV-2 NSP3 and FMRP Interaction
The research uncovered a groundbreaking discovery - a direct interaction between the SARS-CoV-2 NSP3 protein and FMRPs. This interaction, previously unknown, has profound implications for our understanding of how the virus hijacks host cell proteins to bolster its infection capabilities. To demonstrate the significance of this interaction, the study team engineered NSP3 mutant viruses that were incapable of binding to FMRPs. The results were striking: these mutant viruses displayed attenuated replication in vitro and delayed disease onset in vivo. This finding underscores the pivotal role that the NSP3-FMRP interaction plays in the virus's replication and pathogenesis.
Delving into the Molecular Mechanism
To gain deeper insights into this newfound interaction, the study team investigated the molecular details of the binding between NSP3 and FMRPs. They identified a unique peptide motif within NSP3 that directly binds to the central KH domains of FMRPs. This interaction is disrupted by the I304N mutation found in a patient with fragile X syndrome, shedding light on a possible molecular defect associated with this disorder.
The NSP3-FMRP binding disrupts the interaction between FMRPs and UBAP2L, a critical component of stress granules. The direct competition between NSP3 and UBAP2L for binding to FMRPs prevents the incorporation of FMRPs into stress granules. This revelation provides novel insights into how SARS-CoV-2 manipulates host cell proteins to enhance its own replication while modulating stress granule function.
Implications and Future Directions
The discovery of the NSP3-FMRP interaction opens up new avenues of research and potential therapeutic strategies. NSP3's role in disrupting the UBAP2L-FMRP interaction specifically targets stress granule assembly, highlighting the virus's ability to fine-tune its manipulation of host defenses. While the impact of NSP3 mutants on virus replication and disease onset may appear modest, it is crucial to recognize that NSP3 could be instrumental in blunting certain aspects of stress granule formation, particularly during early infection stages, until the N protein takes over to completely disrupt the host process. This mechanism may have a more significant impact in vulnerable individuals, such as the elderly or those with compromised immune systems.
Stress granule formation is an intricately complex process with numerous components and cell-type-specific variations. Stress granules also play critical roles in host responses to various stressors, including viral infections. Therefore, it is not surprising that viruses have evolved multiple strategies to disrupt and co-opt stress granule elements to facilitate their own replication. The importance of stress granules in the host's antiviral response is highlighted by the genetic resources that SARS-CoV-2 dedicates to antagonize them.
The N protein uses its specific motif to prevent interactions with G3BP1/2, thereby disrupting stress granule formation. In this study, NSP3 emerges as an additional stress granule antagonist within the Sarbecovirus family, targeting FMRP integration via UBAP2L. Notably, the peptide sequence required for NSP3's action extends beyond the simple motif found in the N protein. This intriguing finding suggests that the KH domains of FMRP may have binding specificity for certain RNAs, the regulation of which may be affected by NSP3. Given that NSP3 and N protein are in close proximity due to their respective interactions with FMRPs and G3BP1/2, they could work in concert to efficiently disassemble stress granules at sites where nascent viral RNA emerges from double-membrane vesicles, facilitated by NSP3-formed pores.
This intricate viral complex has far-reaching implications for our understanding of SARS-CoV-2 infection and pathogenesis. Furthermore, it opens up exciting possibilities for therapeutic interventions, such as targeting these interfaces to develop direct antiviral drugs and live-attenuated vaccine strategies. However, it's important to acknowledge that interactions with host proteins also raise the possibility of toxicity and off-target effects, necessitating cautious exploration.
In addition to its implications for viral infection, this research offers tantalizing hints about potential molecular defects associated with fragile X syndrome. Emerging data suggest that brain development is intricately linked to stress granule function, but a more detailed mechanistic understanding is required to unravel this complex relationship fully. The detailed molecular insights into the FMRP-UBAP2L complex provided in this study offer a promising avenue to explain its role in brain development and how this role may be impacted by disease mutations like FMR1 I304N.
Moreover, this research raises intriguing questions about the interplay between antiviral defense mechanisms and conditions like fragile X syndrome, or mutations in stress granule components, and how these factors might influence the trajectory of COVID-19 in affected individuals. Further investigations in this direction could uncover novel insights into both viral pathogenesis and neurological disorders.
Conclusion
The study has unveiled a remarkable discovery - the interaction between SARS-CoV-2's NSP3 protein and FMRPs. This interaction not only sheds light on how the virus hijacks host proteins for efficient infection and modulates stress granules but also offers potential insights into the molecular basis of fragile X syndrome. It underscores the complexity of viral-host interactions and the ingenious strategies employed by viruses to manipulate host defenses. This newfound knowledge opens doors to novel therapeutic approaches and paves the way for a deeper understanding of both viral infection and neurological disorders.
The study findings were published on a preprint server and is currently being peer reviewed.
https://www.biorxiv.org/content/10.1101/2023.09.01.555899v1
For the latest
COVID-19 News, keep on logging to Thailand Medical News.
