COVID-19 Immunology: Study Uncovers Role Of T Cell Receptor Signatures In COVID-19 Severity
COVID-19 Immunology - T Cell Receptor Signatures - COVID-19 Severity Jul 22, 2023 2 years, 9 months, 3 weeks, 5 days, 11 hours, 49 minutes ago
While most COVID-19 patients exhibited reduced TCR clone diversity, those with severe disease had a significant increase in the overall abundance of dominant T cell clones, indicating clonal expansion and enrichment of specific TCR clones
COVID-19 Immunology: The COVID-19 pandemic, caused by the novel coronavirus SARS-CoV-2, has left an indelible mark on humanity, claiming millions of lives and altering the course of history. As we grapple with its ongoing impact, researchers from prestigious institutions worldwide are engaged in a tireless quest to understand the virus's complexities and the immune responses it elicits. Among these endeavors, a groundbreaking study conducted by a collaboration of renowned Chinese institutions sought to unravel the enigma of T cell receptor β (TCR β) repertoires in COVID-19 patients, aiming to uncover critical insights into disease severity and pathogenesis.
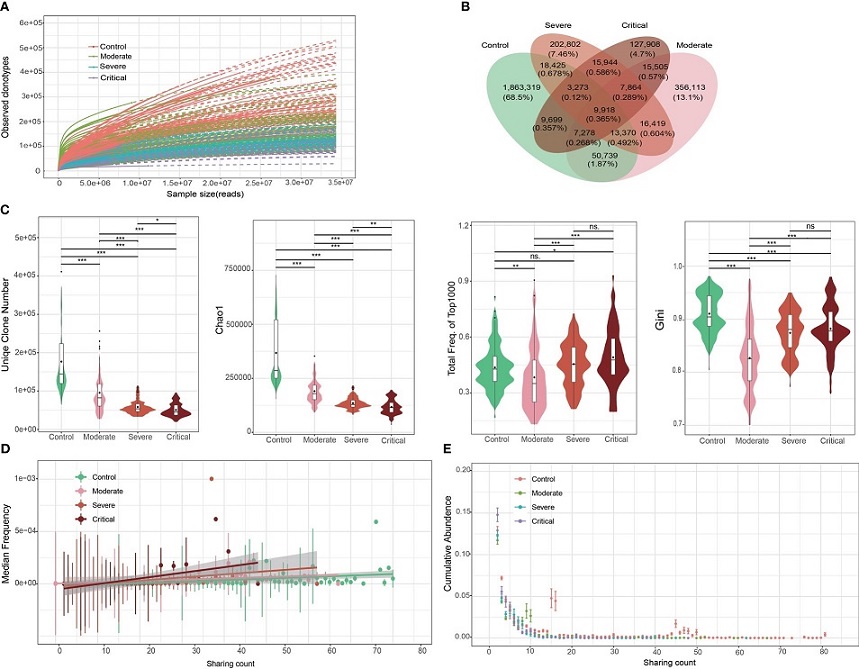 Overall characteristics of the T cell receptor (TCR) β repertoires in patients with COVID-19 and healthy controls (HCs). (A) Saturation analysis of each group. Sequencing data were randomly selected to determine the number of clonotypes detected and extrapolated to the size of the largest samples. Solid and dashed lines represent the interpolated and extrapolated regions of the rarefaction curves, respectively, and points indicate the exact sample size and diversity. (B) Overlap between each group of public clones. (C) Obvious clonal expansion from the aspects of TCR β repertoire diversity in COVID-19 development. Horizontal line is the median, and dot is the average value in the boxplot. (D) Trend of changes in the average abundance of clones under different detection sample numbers for each group. (E) Trend of changes between the total abundance of clones under different detection sample numbers for each group.
Overall characteristics of the T cell receptor (TCR) β repertoires in patients with COVID-19 and healthy controls (HCs). (A) Saturation analysis of each group. Sequencing data were randomly selected to determine the number of clonotypes detected and extrapolated to the size of the largest samples. Solid and dashed lines represent the interpolated and extrapolated regions of the rarefaction curves, respectively, and points indicate the exact sample size and diversity. (B) Overlap between each group of public clones. (C) Obvious clonal expansion from the aspects of TCR β repertoire diversity in COVID-19 development. Horizontal line is the median, and dot is the average value in the boxplot. (D) Trend of changes in the average abundance of clones under different detection sample numbers for each group. (E) Trend of changes between the total abundance of clones under different detection sample numbers for each group.
In this fascinating exploration, the study team delved into the dynamic world of T cells, essential players in our adaptive immune response against viral infections.
The
COVID-19 Immunology researchers aimed to comprehend the delicate balance between effective protection and potentially harmful immune reactions that can drive COVID-19's progression. To achieve this, they employed high-throughput TCR β repertoire sequencing, enabling them to analyze the TCR profiles in the blood of 192 COVID-19 patients, each experiencing various degrees of disease severity, and compared them with 81 healthy individuals.
The study's findings revealed a compelling narrative.
COVID-19 patients exhibited reduced TCR clone diversity, suggesting a more focused and restricted T cell response, likely targeting specific SARS-CoV-2 antigens.
Intriguingly, the team observed a significant increase in the overall abundance of dominant T cell clones in patients with severe disease, indicating clonal expansion and enrichment of specific TCR clones. These TCR β repertoire signatures exhibited the potential to serve as informative biomarkers for monitoring disease progression and severity, offering a promising avenue for diagno
stic applications.
Moreover, the researchers identified significant differences in the genomic rearrangement of variable (V), joining (J), and VJ pairings between the patient groups, highlighting the intricate nuances of T cell immunity in COVID-19.
Armed with this knowledge, the study team harnessed the power of machine learning algorithms to accurately differentiate COVID-19 patients from healthy controls. The team achieved near-perfect accuracy in classifying patients based on TCR β repertoire signatures, underscoring the potential of this approach as a diagnostic tool.
Delving deeper into their exploration, the researchers uncovered a fascinating aspect of TCR repertoires in COVID-19 patients - their heterogeneity. Contrary to the expectation that shared TCR clones would dominate, they discovered a myriad of private TCR clones among patients, showcasing the uniqueness and complexity of individual immune responses. Such heterogeneity suggests that TCR repertoires might hold valuable information for predicting disease severity and aiding in immunodiagnosis.
The study's significance extended beyond COVID-19 itself, as the researchers observed an intriguing enrichment of T cells targeting other viruses, such as Epstein-Barr virus (EBV) and human T-lymphotropic virus 1 (HTLV-1), in COVID-19 patients. This raises questions about potential viral coinfections, molecular mimicry, and cross-activation of T cells, further complicating the already intricate immune responses during COVID-19.
These groundbreaking findings have far-reaching implications. They offer valuable insights into the T-cell immune response and clonal dynamics during COVID-19, deepening our understanding of disease pathogenesis.
The study's emphasis on TCR β repertoires as indicators of disease severity opens new avenues for targeted interventions, personalized medicine, and improved prognostication.
More extensive and diverse cohorts, multi-omics methodologies, and investigations into the interplay between HLA molecules and TCRs promise to unravel even more secrets about the complexities of immune responses to SARS-CoV-2.
In conclusion, the study's findings illuminate the extraordinary world of T cell immunity in COVID-19, revealing TCR β repertoire signatures as potential windows into disease severity and progression.
The study findings were published in the peer reviewed journal: Frontiers In Immunology.
https://www.frontiersin.org/articles/10.3389/fimmu.2023.1190844/full
For the latest on
COVID-19 Immunology, keep on logging to Thailand Medical News.
