Korean Researchers Discover That The Enzyme Trypsin Facilitates SARS-CoV-2 Entry At The Cell Surface
Source: SARS-CoV-2-Research Feb 02, 2022 4 years, 3 months, 4 weeks, 12 hours, 24 minutes ago
SARS-CoV-2-Research: Researchers from Gyeongsang National University-South Korea and the Gyungbuk Veterinary Service Laboratory-South Korea, have in a new study discovered that the enzyme trypsin facilitates SARS-CoV-2 entry at the cell surface.
The coronaviruses typically infect cells by cytoplasmic or endosomal membrane fusion, driven by the spike (S) protein, which must be primed by proteolytic cleavage at the S1/S2 furin cleavage site (FCS) and the S2′ site by cellular proteases.
It has been found that exogenous trypsin serves as a medium additive that facilitates isolation and propagation of several coronaviruses in vitro.
The
SARS-CoV-2-Research team showed that trypsin enhances severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in cultured cells and that SARS-CoV-2 enters cells via either a non-endosomal or an endosomal fusion pathway, depending on the presence of trypsin.
Interestingly, trypsin enabled viral entry at the cell surface led to a more efficient infection than non-trypsin endosomal entry, suggesting that trypsin production or presence in certain organs may trigger a high level of replication of SARS-CoV-2 and cause severe tissue injury.
Extensive syncytium formation and enhanced growth kinetics were observed only in the presence of exogenous trypsin when cell-adapted SARS-CoV-2 strains were tested.
During 50 serial passages without the addition of trypsin, a specific R685S mutation occurred in the S1/S2 FCS (681PRRAR685) that was completely conserved but accompanied by several mutations in the S2 fusion subunit in the presence of trypsin.
The study findings demonstrate that the S1/S2 FCS is essential for proteolytic priming of the S protein and fusion activity for SARS-CoV-2 entry but not for viral replication.
The study findings can potentially contribute to the improvement of SARS-CoV-2 production for the development of vaccines or antivirals and motivate further investigations into the explicit functions of cell-adaptation-related genetic drift in SARS-CoV-2 pathogenesis.
The study findings were published peer reviewed journal: Archives of Virology.
https://link.springer.com/article/10.1007/s00705-021-05343-0
The SARs coronavirus made its debut in Wuhan-China in late December 2019, and to date infected more than 381 million people globally and caused almost 5.7 million COVID-19 related deaths. (The real figures are expected to be around 4 to 6-fold!)
The SARS-CoV-2 virus is the seventh human coronavirus to be discovered and belongs to the Sarbecovirus subgenus of the Betacoronavirus genus in the Coronaviridae family.
The SARS-CoV-2 coronavirus is a single-stranded positive-sense ribonucleic acid (RNA) virus with a large, enveloped genome. SARS-CoV-2 has 16 nonstructural proteins (nsp1–nsp16), four classical coronaviral structural proteins including the spike (S), envelope (E), membrane (M), and nucleocapsid (N) proteins, as well as many accessory proteins in its genome. During SAR
S-CoV-2 infection, the external S protein anchored to the viral E protein interacts with the angiotensin-converting enzyme 2 (ACE2) receptor to promote viral entry into host cells.
Numerous studies have found that the Vero E6 cell line overexpressing transmembrane protease serine 2 (TMPRSS2) exhibited a greater viral titer than parental cells in an infection experiment with SARS-CoV-2, thereby demonstrating the importance of this protease for SARS-CoV-2 fusion capacity and infectivity.
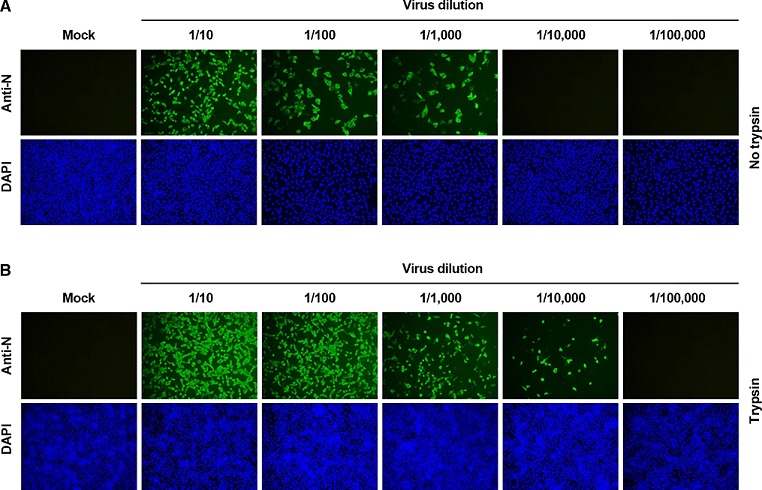 Comparison of SARS-CoV-2 production in the presence and absence of trypsin. Vero E6 cells were mock infected or infected with tenfold serially diluted SARS-CoV-2 and further cultivated in the presence (A) or absence (B) of trypsin. The virus-infected cells were fixed at 24 hpi and incubated with an MAb against the SARS-CoV-2 N protein, followed by incubation with Alexa green–conjugated goat anti-mouse secondary antibody (top panels). The cells were then counterstained with DAPI (bottom panels) and examined under a fluorescence microscope at 200× magnification.
Comparison of SARS-CoV-2 production in the presence and absence of trypsin. Vero E6 cells were mock infected or infected with tenfold serially diluted SARS-CoV-2 and further cultivated in the presence (A) or absence (B) of trypsin. The virus-infected cells were fixed at 24 hpi and incubated with an MAb against the SARS-CoV-2 N protein, followed by incubation with Alexa green–conjugated goat anti-mouse secondary antibody (top panels). The cells were then counterstained with DAPI (bottom panels) and examined under a fluorescence microscope at 200× magnification.
However, trypsin is another protease that has been found to aid in the fusion and infection of human and animal coronaviruses and is required for the multiplication of swine enteric coronaviruses such as porcine endemic diarrhea virus (PEDV) and porcine deltacoronavirus (PDCoV) in vitro.
In this new study in South Korea, exogenous trypsin treatment was found to boost SARS-CoV-2 infection in Vero E6 cells and promote viral multiplication in cultured cells.
It was found that none of the trypsin dosages tested in this study caused detectable cell death, according to the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) cell viability assay. At 24 hours post-infection (HPI), the study team used virus titration to measure virus production.
Irrespective of whether trypsin was present or not before infection, viral yields were constant, with a mean titer range of 106.53–106.73 50% tissue culture infectious dose (TCID50)/ml.
The strength of the cytopathic effect (CPE) was also monitored for SARS-CoV-2 replication, which was validated by immunofluorescence assay (IFA) employing an anti-N-protein monoclonal antibody (MAb) at 24 HPI.
Interestingly when compared to the untreated control, cells pre-treated with trypsin prior to viral infection showed no effect on SARS-CoV-2 infectivity at any of the test concentrations.
Also, introducing exogenous trypsin during the one-hour inoculation periods had no effect on SARS-CoV-2 propagation in cell culture. Thus, trypsin therapy before or during infection had no effect on SARS-CoV-2 replication in cells.
However, in the presence of 5 µg/ml of trypsin, the greatest viral titer of 107.52 TCID50/ml was achieved, which was one log greater than the control mean titer. When SARS-CoV-2-infected cells were cultured in the presence of trypsin, CPE and IFA analyses revealed that virus infectivity increased.
In another experiment by the team, Vero E6 cells were inoculated with ten-fold serially diluted SARS-CoV-2 and kept in the presence or absence of trypsin, whereas IFA was used to visualize infected cells.
The study team discovered that when Vero E6 cells were propagated with trypsin, the number of infected cells was more than ten-fold higher than when the cells were propagated without trypsin.
The study team then investigated whether trypsin helps SARS-CoV-2 enter cultured cells. To this end, the team employed a proteinase K infection assay to measure the rate of virus attachment and penetration by quantifying bound and internalized virus particles, respectively, to set the parameters for the viral internalization assay used to monitor SARS-CoV-2 entrance.
The vero E6 cells were then infected with SARS-CoV-2 for one hour at 4°C and then treated with proteinase K at various doses for 45 minutes at 4°C to test the efficiency of proteinase K therapy in eliminating bound viruses. The SARS-CoV-2 genome copy number was determined by measuring the amount of virus attached to the cells using a quantitative reverse-transcriptase polymerase chain reaction (qRT-PCR) assay.
The study findings showed that proteinase K treatment was found to reduce the number of virions bound to the cell surface significantly, thereby implying that SARS-CoV-2 that is bound to cells is effectively eliminated by proteinase K treatment.
The study findings demonstrate that trypsin improves SARS-CoV-2 multiplication in cultivated cells and aids viral entrance by facilitating a direct fusion process at the cell surface.
The study findings also suggest that depending on the presence of trypsin, SARS-CoV-2 may adopt distinct paths to enter cells.
It was also noted that the non-endosomal entrance of SARS-CoV-2 at the cell surface triggered by trypsin facilitated more effective infection than the endosomal pathway in the absence of trypsin.
For the latest
SARS-CoV-2-Research, keep on logging to Thailand Medical News.

