Study Reveals Alarming Rate of Protein-Coding Changes In SARS-CoV-2 Compared To Other Endemic Viruses
COVID-19 News - SARS-CoV-2 - Other Endemic Viruses May 23, 2023 2 years, 11 months, 1 week, 5 days, 19 hours, 34 minutes ago
COVID-19 News: A recent study conducted by researchers from the Vaccine and Infectious Disease Division at the Fred Hutchinson Cancer Center and the Howard Hughes Medical Institute in Seattle-USA has shed light on the concerning evolutionary dynamics of SARS-CoV-2, the virus responsible for the COVID-19 pandemic.
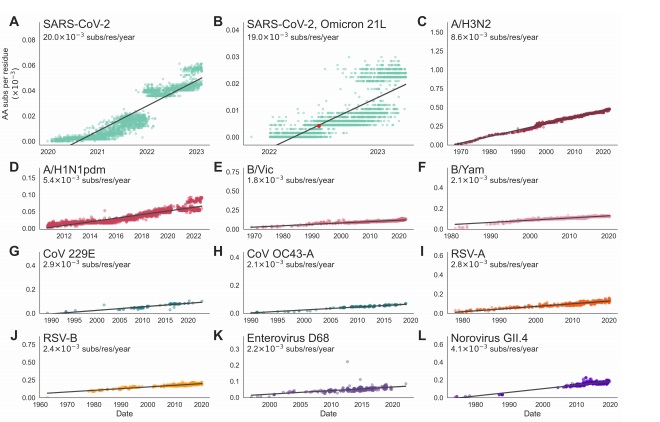 Rates of amino acid substitution in the receptor-binding protein of SARS-CoV-2 and 10 antigenically-evolving endemic viruses. The rate of amino acid substitution in the receptor-bindingprotein of A) SARS-CoV-2, B) SARS-CoV-2 Omicron clade 21L, C) influenza A/H3N2, D) influenza
Rates of amino acid substitution in the receptor-binding protein of SARS-CoV-2 and 10 antigenically-evolving endemic viruses. The rate of amino acid substitution in the receptor-bindingprotein of A) SARS-CoV-2, B) SARS-CoV-2 Omicron clade 21L, C) influenza A/H3N2, D) influenza
A/H1N1pdm, E) influenza B/Vic, F) influenza B/Yam, G) coronavirus 229E, H) coronavirus OC43-A, I)RSV-A, J) RSV-B, K) enterovirus D68, L) norovirus GII.4. Rates are computed as the slope of a linear regression fitting a comparison of amino acid substitutions versus time, and are found using a phylogeny. Each tip on the tree is plotted by its sampling date and the number of amino acid substitutions that accumulated between the root and the tip (normalized by the length of the coding region, in residues). Aspect ratios in each panel are fixed so that regression slopes are visually comparable across panels.
The study compared the rates of adaptive evolution in SARS-CoV-2 with those of 28 endemic human viruses, spanning various viral families and transmission modes. The study findings indicate that SARS-CoV-2 is accumulating protein-coding changes at significantly faster rates than other endemic viruses, raising important questions about the virus's ability to evade immunity and the potential implications for vaccine development.
As covered by numerous
COVID-19 News coverages, the SARS-CoV-2 virus especially the Omicron variants and XBB sub-lineages are evolving at an unprecedented rate and spawning newer sub-lineages with a variety of unique mutations as it evolves to evade the various forms of immunity. At present we have more than 207 different XBb sub-lineages in circulation globally with many more emerging continuously.
https://www.thailandmedical.news/news/more-than-207-sars-cov-2-xbb-sublineages-in-circulation-globally-in-last-30-days-and-more-to-come-covid-19-public-health-emergency-over
Understanding Antigenic Evolution
Viruses possess a remarkable ability to rapidly evolve in response to changing environments, which allows them to persist and thrive. Antigenic evolution, in particular, enables viruses to evade recognition by neutralizing antibodies generated through previous infection or vaccination. This phenomenon has been extensively observed in seasonal influenza viruses, necessitating frequent updates of the influenza vaccine to keep up with the evolving strains. Antigenic evolution also occurs in other viruses such as coronaviruses and noroviruses.
The Study's Approach
To expand the knowledge of antigenic evolution in endemic human viruses, the
study team analyzed adaptive evolution across the genomes of 28 viruses. They focused on the protein-coding changes in the viral genome, particularly in the receptor-binding proteins responsible for antibody recognition and neutralization.
The rates of adaptive evolution were estimated using a quantitative method based on genetic sequences of viral isolates collected over time. This approach allowed for comparisons within and between viruses to identify those undergoing antigenic evolution.
Results and Implications
The study revealed that ten viruses, including seasonal influenza A and B viruses, norovirus, respiratory syncytial virus, and certain coronaviruses, exhibited elevated rates of adaptation in their receptor-binding proteins. These findings suggest ongoing antigenic evolution in these viruses, enabling them to cause repeat infections and escape vaccine-mediated immunity.
Notably, the study demonstrated that SARS-CoV-2 is accumulating protein-coding changes at significantly faster rates than these endemic viruses. In fact, the rate of amino acid substitution in SARS-CoV-2's S1 subunit was found to be roughly 2-2.5 times higher than that of the prototypical rapidly evolving influenza A/H3N2 strain.
This raises concerns about the virus's potential to sustain high rates of evolution in the future, particularly as complex immune histories towards SARS-CoV-2 continue to develop in the population.
While the exact factors driving antigenic evolution in different viruses remain unclear, the study highlights the need for a comprehensive understanding of ongoing viral evolution. By expanding our knowledge of endemic viruses, we can better prepare for future pandemics and enhance our efforts in designing effective vaccines and therapeutics.
The Way Forward
The research team behind this study have developed an online resource, the Atlas of Viral Adaptive Evolution, which provides interactive plots and phylogenies to further explore the results and compare the evolution of different viruses.
https://blab.github.io/atlas-of-viral-adaptation/
Additionally, ongoing efforts to monitor and sequence human pathogens will contribute to the accumulation of longitudinal data, enabling the inclusion of more viruses in future studies.
Conclusion
The findings of this study emphasize the rapid adaptive evolution of SARS-CoV-2 compared to other endemic human viruses. The accelerated accumulation of protein-coding changes in the virus raises concerns about its ability to evade immunity and underscores the need for ongoing surveillance, vaccine updates, and therapeutic development. By expanding our understanding of viral evolution, we can better respond to emerging pathogens and mitigate the impact of future pandemics.
The study findings were published on a preprint server and are currently being peer reviewed.
https://www.biorxiv.org/content/10.1101/2023.05.19.541367v1
For the latest
COVID-19 News, keep on logging to Thailand Medical News.
