BREAKING! US Scientists Find That COVID-19 Leaves Lasting Epigenetic Footprint That Results In Long-Term Immune Dysregulation!
COVID-19-News-Lasting-Epigenetic-Footprint-Long-Term-Immune-Dysregulation May 29, 2023 3 years, 3 days, 15 hours, 52 minutes ago
COVID-19 News: In the midst of the COVID-19 pandemic that has affected millions worldwide, scientists have been tirelessly working to understand the long-term consequences of the disease. Despite survivors experiencing persistent symptoms, the underlying mechanisms responsible for these lingering effects have remained elusive. However, a recent research conducted by Albany Medical College in New York, USA, and the University of Wisconsin School of Medicine and Public Health in Madison, USA, has shed new worrisome findings on the subject.
 The Research
The Research
The study team focused on epigenetic mechanisms, which control gene expression without altering the underlying DNA sequence. Specifically, they examined DNA methylation, a process that can modulate gene activity and is influenced by environmental factors. While previous studies had shown abnormal DNA methylation in patients recovering from severe SARS-CoV-2 infection, this new study aimed to explore the entire genome using cutting-edge sequencing technologies.
Thailand Medical News had already published previous
COVID-19 News reports that showed SARS-CoV-2 infections can lead to epigenetic changes in the infected human host.
https://www.thailandmedical.news/news/long-covid-news-sars-cov-2-infections-causes-epigenetic-changes-that-contributes-to-health-issues-seen-in-long-covid
https://www.thailandmedical.news/news/south-african-study-finds-that-covid-19-leads-to-epigenetic-changes-in-host-immune-genes-which-can-be-prevented-or-reversed-by-vitamin-d
https://www.thailandmedical.news/news/international-study-finds-that-sars-cov-2-causes-epigenetic-changes-and-transcriptomic-reprogramming-in-monocytes-of-covid-19-patients
https://www.thailandmedical.news/news/breaking-swedish-study-shows-that-sars-cov-2-causes-epigenetic-changes-to-various-genes-in-human-host
https://www.thailandmedical.news/news/breaking-u-s-study-finds-that-sars-cov-2-infections-and-mrna-jabs-alter-dna-methylation-states-of-immune-cells-and-epigenetic-clocks-of-humans
-causes-epigenetic-changes-to-innate-immune-cells-and-their-progenitors,-contributing-to-long-covid">https://www.thailandmedical.news/news/new-study-by-cornell-university-shows-sars-cov-2-causes-epigenetic-changes-to-innate-immune-cells-and-their-progenitors,-contributing-to-long-covid
Worrisome Research Findings
The study team compared samples from patients admitted for COVID-19 with samples taken one year after their hospital discharge. They also analyzed samples from healthy individuals who had never been diagnosed with COVID-19. The analysis revealed distinct DNA methylation patterns in patients with acute infection compared to healthy volunteers.
Remarkably, even one year after recovery, these patients exhibited persistent alterations in DNA methylation.
The Impact of Epigenetic Dysregulation
The persistent changes in DNA methylation were primarily located in regions near genes involved in immune response pathways. Importantly, the regions that did not return to baseline methylation levels after one year correlated with immune dysregulation and lymphocyte activation, which are known to be associated with prolonged COVID-19 symptoms. This suggests that altered DNA methylation may contribute to the long-lasting effects experienced by many COVID-19 survivors.
Uncovering the Whole Genome
What sets this study apart is the use of whole-genome methylation sequencing, which allowed the study team to examine approximately 25 million sites in the genome. This comprehensive approach enabled the identification of additional regions with significant dysregulation that were not covered by previous microarray analyses. By employing this unbiased technique, the researchers unveiled the extent of the methylation changes induced by acute COVID-19 infection.
Future Implications
These study findings open up new avenues of research and underscore the importance of investigating epigenetic changes in COVID-19 survivors.
By conducting larger studies with diverse populations, scientists hope to further pinpoint the specific regions and gene transcripts involved in long-term COVID-19 sequelae. This knowledge could pave the way for targeted therapies to alleviate the persistent symptoms experienced by survivors.
Study Limitations and Future Directions
While this study provides valuable insights, there are limitations that should be addressed in future research. The control group consisted of healthy volunteers, potentially introducing interindividual differences that may not be directly related to the disease stage. Additionally, the relatively small sample size may have affected the statistical significance of certain findings. Further studies should also consider the influence of comorbidities and explore specific clinical deficits associated with sustained post-COVID-19 effects.
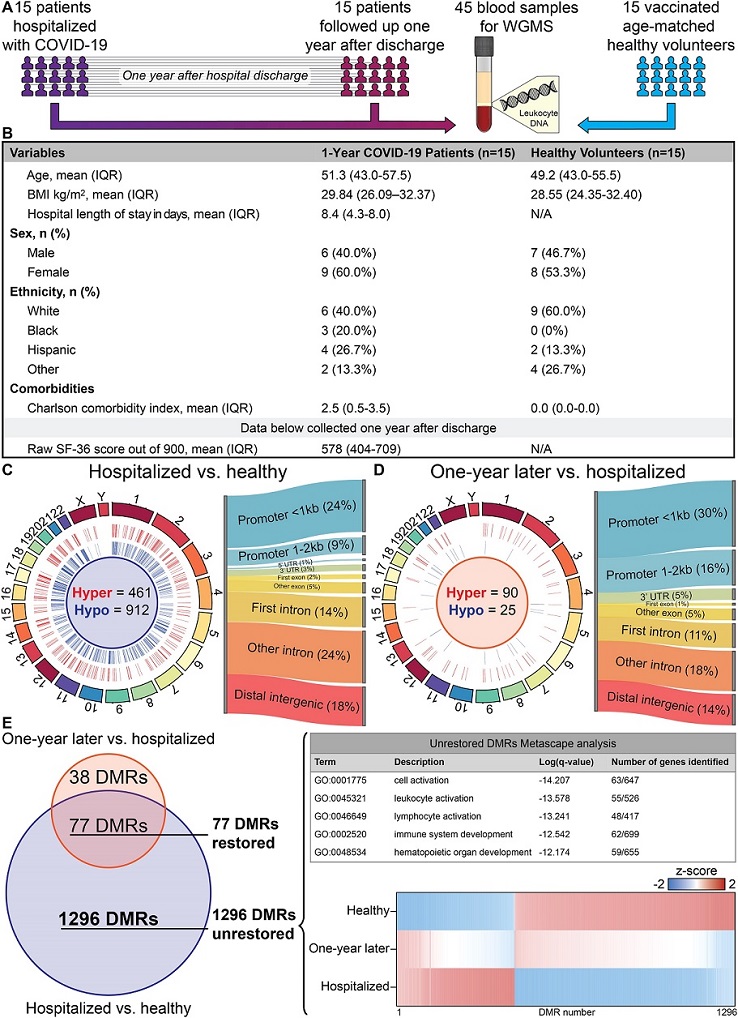 WGMS differential analysis of age-matched healthy volunteers, 15 acutely ill patients hospitalized with coronavirus disease (COVID-19), and the same 15 patients recalled 1 year after hospital discharge. (A) Schematic of study design and sample collections. See text for details. (B) Clinical characteristics of participants. The raw Short Form Health Survey involves 36 questions that are divided into nine domains. Each domain has a maximal score of 100% based on the participants’ answers, so the optimal score is 900. (C) Circos plot showing the genomic distribution of differentially methylated regions (DMRs) across the human genome. Each chromosome is shown as a different color in the ring. Relative chromosome size is denoted by arc bar length. Hypermethylated DMRs are in red, and hypomethylated regions are in blue. This comparison identified 1,373 DMRs that were differentially methylated within days of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. The Sankey diagram shows the percent distribution of DMRs to standard genomic features. A preponderance of DMRs are observed in gene promoter regions. (D) Circos plot (see C for description) of paired samples 1 year after hospital discharge versus during hospitalization with severe COVID-19. The paired comparison identified 115 DMRs differentially methylated 1 year after hospital discharge. The Sankey diagram shows the percent distribution of DMRs to standard genomic features. A preponderance of DMRs are observed in gene promoter regions. (E) A Venn diagram overlapping DMRs from the comparisons made in C and D identified 77 DMRs that return to baseline levels of methylation after acute infection and 1,296 covid-induced DMRs that do not return to baseline in patients after 1 year of follow-up. Metascape analysis of the genes with DMRs that do not return to baseline in patients recalled 1 year after discharge with terms that correspond to cell activation and immune response. A heat map of the methylation z-score of the 1,296 unrestored DMRs for the three populations shows DMRs that do not recover to baseline DNA methylation levels 1 year after acute COVID-19 infection. IQR = interquartile range; N/A = not applicable; UTR = untranslated region; WGMS = whole-genome methylation sequencing
Conclusion
WGMS differential analysis of age-matched healthy volunteers, 15 acutely ill patients hospitalized with coronavirus disease (COVID-19), and the same 15 patients recalled 1 year after hospital discharge. (A) Schematic of study design and sample collections. See text for details. (B) Clinical characteristics of participants. The raw Short Form Health Survey involves 36 questions that are divided into nine domains. Each domain has a maximal score of 100% based on the participants’ answers, so the optimal score is 900. (C) Circos plot showing the genomic distribution of differentially methylated regions (DMRs) across the human genome. Each chromosome is shown as a different color in the ring. Relative chromosome size is denoted by arc bar length. Hypermethylated DMRs are in red, and hypomethylated regions are in blue. This comparison identified 1,373 DMRs that were differentially methylated within days of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. The Sankey diagram shows the percent distribution of DMRs to standard genomic features. A preponderance of DMRs are observed in gene promoter regions. (D) Circos plot (see C for description) of paired samples 1 year after hospital discharge versus during hospitalization with severe COVID-19. The paired comparison identified 115 DMRs differentially methylated 1 year after hospital discharge. The Sankey diagram shows the percent distribution of DMRs to standard genomic features. A preponderance of DMRs are observed in gene promoter regions. (E) A Venn diagram overlapping DMRs from the comparisons made in C and D identified 77 DMRs that return to baseline levels of methylation after acute infection and 1,296 covid-induced DMRs that do not return to baseline in patients after 1 year of follow-up. Metascape analysis of the genes with DMRs that do not return to baseline in patients recalled 1 year after discharge with terms that correspond to cell activation and immune response. A heat map of the methylation z-score of the 1,296 unrestored DMRs for the three populations shows DMRs that do not recover to baseline DNA methylation levels 1 year after acute COVID-19 infection. IQR = interquartile range; N/A = not applicable; UTR = untranslated region; WGMS = whole-genome methylation sequencing
Conclusion
The worrisome study findings reveal that COVID-19 leaves a lasting epigenetic footprint in survivors. By analyzing the entire genome, the study team have discovered persistent alterations in DNA methylation one year after recovery.
These study findings emphasize the need for continued investigation into the molecular mechanisms underlying long-term COVID-19 symptoms. By unraveling the complexities of epigenetic dysregulation, scientists can work towards developing targeted interventions to improve the lives of those affected by this unprecedented global health crisis.
The study findings were published as a correspondence in the peer reviewed American Journal of Respiratory Cell and Molecular Biology.
https://www.atsjournals.org/doi/full/10.1165/rcmb.2022-0433LE
For the latest
COVID-19 News, keep on logging to Thailand Medical News.

