Study Identifies Seven Hub Genes IFI44, IFI44L, MX1, OAS3, USP18, IFI27 And ISG15 As Potential Therapeutic Targets For SARS-CoV-2 Omicron Infections
Thailand Medical News Team Aug 18, 2023 2 years, 9 months, 3 days, 18 hours, 14 minutes ago
COVID-19 News: The COVID-19 pandemic, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has brought forth an unprecedented global health crisis. As the virus continues to evolve, its various variants present a complex spectrum of clinical outcomes, ranging from asymptomatic cases to severe respiratory illnesses. The emergence of the SARS-CoV-2 Omicron variant has further accentuated the need to understand the underlying mechanisms driving symptom development and to identify potential therapeutic targets. In this pursuit, researchers at the First Affiliated Hospital of Guangzhou Medical University-China and the Guangzhou Laboratory-China conducted a comprehensive study to unravel the role of hub genes in SARS-CoV-2 Omicron infection, aiming to shed light on its clinical manifestations and potential treatment strategies.
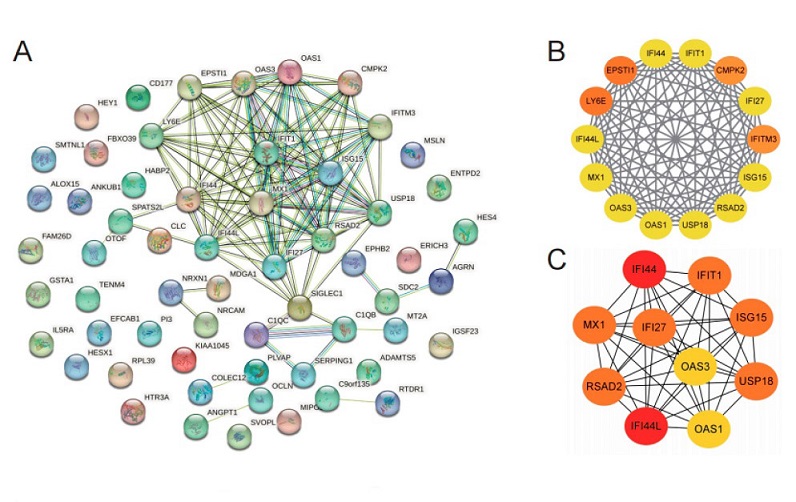 Identification and validation of hub genes. STRING (A); MCODE hub genes (B); cytohubba-MCC hub genes (C).
Exploring the Omicron Variant's Impact
Identification and validation of hub genes. STRING (A); MCODE hub genes (B); cytohubba-MCC hub genes (C).
Exploring the Omicron Variant's Impact
The SARS-CoV-2 Omicron variant, characterized by its rapid transmission and altered spike protein, has been associated with milder clinical manifestations in some regions, marked by lower hospitalization rates and shorter lengths of stay as covred in various studies and
COVID-19 News reports. However, despite this apparent reduction in severity, the global healthcare system continues to be strained, emphasizing the importance of understanding the molecular mechanisms underlying the variant's symptomatic presentation. The study conducted by the researchers sought to bridge this knowledge gap by analyzing transcriptome datasets from peripheral blood mononuclear cells (PBMCs) of COVID-19 patients infected with the Omicron variant and healthy volunteers.
Identifying Hub Genes as Biomarkers
The research team utilized a bioinformatics approach to identify hub genes associated with the Omicron variant. Through the analysis of gene expression datasets and the construction of protein-protein interaction networks, they pinpointed ten hub genes that showed promise as potential biomarkers. To validate these findings, an independent dataset of COVID-19 patients' PBMCs was employed, consistently confirming the expression patterns of the identified hub genes and their diagnostic value. Further analysis of datasets representing various stages of infection revealed seven key hub genes - IFI44, IFI44L, MX1, OAS3, USP18, IFI27, and ISG15 - that correlated with symptom onset and recovery from COVID-19.
Understanding Type I Interferon-Related Pathways
Type I interferons (IFN-I) have emerged as pivotal players in the antiviral response against SARS-CoV-2. These signaling molecules activate a cascade of interferon-stimulated genes (ISGs) through the JAK-STAT pathway, contributing to viral clearance and immune modulation. The researchers noted that the identified hub genes were predominantly associated with IFN-I-related pathways. MX1, OAS3, USP18, IFI27, and ISG15 were particularly noteworthy, as they have been shown to regulate antiviral responses. The study's findings provided
valuable insights into the intricate interplay between IFN-I pathways and the progression of COVID-19 symptoms, offering potential avenues for therapeutic intervention.
Immune Cell Analysis and Clinical Correlations
In-depth immune cell analysis using the "CIBERSORTx" tool revealed altered proportions of immune cell types in COVID-19 patients, with a notable increase in monocytes and a decrease in certain subsets of T cells. This observation was consistent with clinical data from symptomatic COVID-19 patients with Omicron infection, further substantiating the link between elevated monocyte levels and symptomatic development. These findings not only highlighted the importance of immune cell dynamics in COVID-19 pathogenesis but also strengthened the study's overall conclusions.
Potential Therapeutic Targets and Implications
The identification of the seven hub genes as potential biomarkers for diagnosing and treating Omicron infection holds significant clinical implications. By targeting these hub genes, researchers have the opportunity to develop targeted therapeutic interventions that modulate IFN-I-related pathways, potentially enhancing viral clearance and mitigating symptom severity. The study also explored potential therapeutic agents using the DSigDB database, with acetohexamide emerging as a promising candidate for COVID-19 treatment. The validation and further investigation of these potential therapies represent a promising avenue for future research.
Conclusion
In conclusion, the study provides a comprehensive understanding of the molecular mechanisms underlying SARS-CoV-2 Omicron infection. By identifying and validating hub genes associated with IFN-I-related pathways, the study not only sheds light on the development and recovery of COVID-19 symptoms but also offers potential therapeutic targets for mitigating the impact of the disease.
The implications of this research extend beyond the Omicron variant, offering valuable insights into the broader landscape of COVID-19 pathogenesis and treatment strategies.
The study findings were published in the peer reviewed journal: Microorganism.
https://www.mdpi.com/2076-2607/11/8/2101
For the latest
COVID-19 News, keep on logging to Thailand Medical News.
