Study Discovers That SARS-CoV-2 Mpro Protein Disarms Some Of The Human Immune Responses By Cleaving Nemo, An Immune Signaling Protein!
Source: Medical News - SARS-CoV-2 Mpro Cleaves Human Proteins Sep 09, 2022 3 years, 8 months, 2 weeks, 1 day, 14 hours, 18 minutes ago
A new study led by researchers from the SLAC National Accelerator Laboratory at Stanford University and the National Virtual Biotechnology Laboratory at the US Department of Energy-Washington has found the SARS-CoV-2 main protease Mpro also called
3CLpro (3-chymotrypsin like protease ) is able to cleave a key human immune signaling protein called NEMO (NF-κB Essential Modulator) in the process severing certain critical immune pathways and disarming certain immune responses!
More alarmingly, the study indicates that many more other human host proteins can also be cleaved by the Mpro, contributing to either immediate or long-term medical conditions…..many of which we are not even aware of currently!
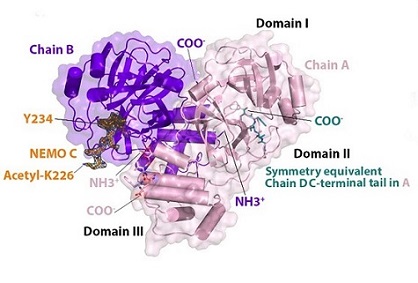 The hNEMO226–234-bound 3CLpro dimer. N- (NH3 + ) and C-termini (COO-) are labeled. hNEMO226–234 (NEMO B - orange) binds into chain B (purple) and is surrounded by an omit map of electron density (1.0 σ contour level and 1.9 Å carving radius). Acetylated Lys226 and Tyr234 at the N- and C-termini of the hNEMO226–234 peptide, respectively, are labeled. The C-terminal tail of chain D (teal) from a crystallographic symmetry equivalent binds into the substrate-binding site of chain A (light pink). Domains I, II and III are labeled.
The hNEMO226–234-bound 3CLpro dimer. N- (NH3 + ) and C-termini (COO-) are labeled. hNEMO226–234 (NEMO B - orange) binds into chain B (purple) and is surrounded by an omit map of electron density (1.0 σ contour level and 1.9 Å carving radius). Acetylated Lys226 and Tyr234 at the N- and C-termini of the hNEMO226–234 peptide, respectively, are labeled. The C-terminal tail of chain D (teal) from a crystallographic symmetry equivalent binds into the substrate-binding site of chain A (light pink). Domains I, II and III are labeled.
The study also involved researchers from the University of Kentucky, Chapman University School of Pharmacy, Oak Ridge National Laboratory and the University of Tennessee.
The
SARS-CoV-2 3C-like protease (3CLpro
) plays a major role in the viral replication and is also essential in viral polyprotein processing.
The NEMO proteins are part of a human immune system known as the NF-κB pathway.
One can can think of NEMO and the NF-κB pathway as if they were a card reader and wiring on the outside of a locked building entrance door. If the wires to the card reader are cut, the door will not open, meaning a person (or an immune system activator, like NEMO) is stuck outside, unable to do whatever they came to do.
Corresponding author, Dr Soichi Wakatsuki, a professor at SLAC and at Stanford University explained to Thailand
Medical News, “Importantly, the NF-κB pathway is a critical part of protective inflammatory responses. When NEMO is cut, human immune responses can't be activated, resulting in various detrimental effects to our body. COVID-19 viral infections could be made worse if Mpro destroys NEMO, helping the virus evade the innate immune responses.”
The study involving vitro assays showed that SARS-CoV-2 3CLpro cleaves NEMO with fine-tuned efficiency.
Subsequent detailed analysis of the 2.50 Å resolution crystal structure of 3CLpro C145S bound to NEMO226–234 revealed subsites that tolerate a range of viral and host substrates through main chain hydrogen bonds while also enforcing specificity using side chain hydrogen bonds and hydrophobic contacts.
Machine learning- and physics-based computational methods predicted variation in key bindin
g residues of 3CLpro-NEMO, helping to explain the high fitness of SARS-CoV-2 in humans.
The study team posit that cleavage of NEMO is an important piece of information to be accounted for, in the pathology of COVID-19.
The study findings were published in the peer reviewed journal: Nature Communications.
https://www.nature.com/articles/s41467-022-32922-9
In the last 30 months, researchers and scientists have studied the SARS-CoV-2 virus in great detail, laying the foundation for developing COVID-19 vaccines and antiviral treatments.
However, this is the first time, scientists at the Department of Energy's SLAC National Accelerator Laboratory have seen one of the virus's most critical interactions, which could help researchers develop more precise treatments.
The study team caught the moment when a virus protein, called Mpro, cuts a human protective protein, known as NEMO, in an infected individual.
Without NEMO, the human immune system is slower to respond to increasing viral loads or new infections. Seeing how Mpro attacks NEMO at the molecular level could inspire new therapeutic approaches.
In order to observe how Mpro cuts NEMO, the study team funneled powerful X-rays from SLAC's Stanford Synchrotron Radiation Lightsource (SSRL) onto crystallized samples of the protein complex. The X-rays struck the protein samples, revealing what Mpro looks like when it dismantles NEMO's primary function of helping our immune system communicate.
Dr Wakatsuki said,"We saw that the virus protein cuts through NEMO as easily as sharp scissors through thin paper. Imagine the bad things that happen when good proteins in our bodies start getting cut into pieces!"
The detailed images from SSRL showed the exact location of NEMO's cut and provided the first structure of SARS-CoV-2 Mpro bound to a human protein.
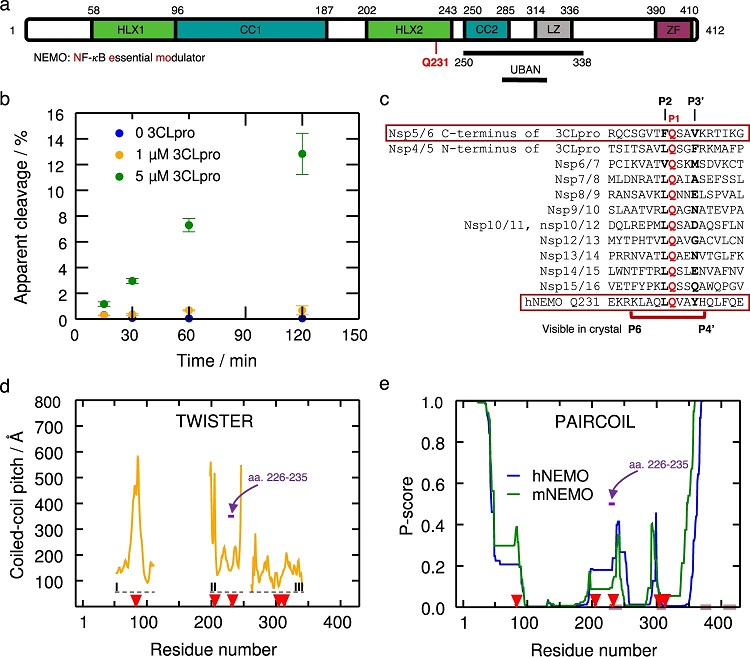 a NEMO includes the α-helical domain 1 (Hlx1), the coiled-coil domain 1 (CC1), the α-helical domain 2 (Hlx2), the coiled-coil domain (CC2), a leucine zipper (LZ) domain, and the C-terminal zinc-finger (ZF). Human NEMO (hNEMO) truncated at site 215 and 247 was used in enzymatic assays with 3CLpro. A recognition site of cleavage is found at Gln231. b Cleavage of hNEMO215–247 at 0.053 μg/μL (~13 μM) by 3CLpro at two concentrations. Reactions were incubated at 25 °C and aliquots were quenched at different times for analysis. The extent of proteolysis was quantified by LC-MS/MS. Apparent % cleavage was calculated by dividing the product peak area by the sum of the substrate and product peak areas. Error bars represent the range of duplicate enzymatic reactions. Statistics have been derived for n = 2 biologically independent experiments. c Multiple sequence alignment of peptide sequences of SARS-CoV-2 polyprotein and hNEMO. P1 site glutamine residues are shown in red. The peptide (P6 to P4’) used in the crystal structure is indicated beneath the sequences. d Coiled-coil pitch per residue computed with TWISTER24 for NEMO in the PDB structures 6MI3 (region I)34, 3CL3 (region II)33, and 6YEK (region III)73 is shown. Dashed lines indicate the regions I-III corresponding to these structures. The cleavage sites Gln83, Gln205, Gln 231, Gln304, and Gln313 are indicated with red arrows. The region corresponding to hNEMO226–235, used in our X-ray structure determination is pointed out (violet arrow). e PAIRCOIL25 prediction of coiled-coil propensity per residue for human and mouse NEMO (mNEMO). Lower P-scores implies greater likelihood of coiled-coil. Potential disordered binding regions predicted by ANCHOR26 are shown in brown. The cleavage sites are depicted as in d. The region corresponding to hNEMO226–235 used in our X-ray structure is depicted as in d.
a NEMO includes the α-helical domain 1 (Hlx1), the coiled-coil domain 1 (CC1), the α-helical domain 2 (Hlx2), the coiled-coil domain (CC2), a leucine zipper (LZ) domain, and the C-terminal zinc-finger (ZF). Human NEMO (hNEMO) truncated at site 215 and 247 was used in enzymatic assays with 3CLpro. A recognition site of cleavage is found at Gln231. b Cleavage of hNEMO215–247 at 0.053 μg/μL (~13 μM) by 3CLpro at two concentrations. Reactions were incubated at 25 °C and aliquots were quenched at different times for analysis. The extent of proteolysis was quantified by LC-MS/MS. Apparent % cleavage was calculated by dividing the product peak area by the sum of the substrate and product peak areas. Error bars represent the range of duplicate enzymatic reactions. Statistics have been derived for n = 2 biologically independent experiments. c Multiple sequence alignment of peptide sequences of SARS-CoV-2 polyprotein and hNEMO. P1 site glutamine residues are shown in red. The peptide (P6 to P4’) used in the crystal structure is indicated beneath the sequences. d Coiled-coil pitch per residue computed with TWISTER24 for NEMO in the PDB structures 6MI3 (region I)34, 3CL3 (region II)33, and 6YEK (region III)73 is shown. Dashed lines indicate the regions I-III corresponding to these structures. The cleavage sites Gln83, Gln205, Gln 231, Gln304, and Gln313 are indicated with red arrows. The region corresponding to hNEMO226–235, used in our X-ray structure determination is pointed out (violet arrow). e PAIRCOIL25 prediction of coiled-coil propensity per residue for human and mouse NEMO (mNEMO). Lower P-scores implies greater likelihood of coiled-coil. Potential disordered binding regions predicted by ANCHOR26 are shown in brown. The cleavage sites are depicted as in d. The region corresponding to hNEMO226–235 used in our X-ray structure is depicted as in d.
SSRL lead scientist and co-author Dr Irimpan Mathews added, "If you can block the sites where Mpro binds to NEMO, you can stop this cut from happening over and over. Stopping Mpro could slow down how fast the virus takes over a body. Solving the crystal structure revealed Mpro's binding sites and was one of the first steps to stopping the protein."
The disarming of the human immune responses and severing of certain critical immune pathways by the virus Mpro proteins cleaving the human Nemo protein could also account for certain manifestations seen in Long COVID.
Interesting, there is also a separate study by researchers from institutions in Germany that found the loss of NEMO by the action of Mpro could lead to damage in certain brain cells, causing neurological symptoms observed in COVID-19 patients!
https://www.nature.com/articles/s41593-021-00926-1
Drugs that targets Mpro proteins ie Mpro inhibitors could help in treating the infection and development of such drugs can be enhanced by this new study findings
SLAC scientist and co-first author Mikhail Ali Hameedi added, "The crystal structures of NEMO and Mpro provide us with the targets to develop treatments that stop these cuts from happening. Although current antiviral drugs can target Mpro, seeing the molecular details of how Mpro attacks NEMO will help us develop new treatments in the future as Mpro mutates."
Strategies to improve antiviral inhibitors is especially important with SARS-CoV-2.
It found that among the all the coronaviruses including the original SARS-CoV and MERS‐CoV viruses, the SARS-CoV-2's Mpro is the most effective at attaching to and cutting NEMO.
The SARS-CoV-2's Mpro grabs NEMO with a tighter grip than its counterparts in other coronaviruses and could be cutting hundreds of other critical proteins in human host cells, such as those associated with blood disorders, the study team warned.
In order to predict how well Mpro binds to NEMO, the study team used the Summit supercomputer at the Oak Ridge Leadership Computing Facility. They combined molecular dynamics simulations with five machine learning models in a novel way and applied quantum chemistry, finding that Mpro likely has the highest binding affinity in SARS-CoV-2 compared to the other primary coronaviruses.
In past studies, these techniques helped scientists narrow down a list of potential antiviral inhibitor drugs.
Co-first author and ORNL scientist Dr Erica Prates added, "With a set of computational approaches, we were able to predict the strongest binding spots between NEMO and Mpro. We think that a high binding affinity at these hot spots helps explain the high fitness of the virus in humans."
Dr Wakatsuki added, “The biomedical industry could use the study to help build better inhibitor drugs and understand how other proteins could be affected by Mpro.”
But he warned, “NEMO is only the tip of the iceberg. We suspect that there are many more human proteins that can be cleaved by the Mpro and even more worrying, with the SARS-CoV-2 evolving and mutating rapidly resulting in the emergence in so many new variants and subvariants, we have no clue what is actually happening inside the human body over time and with new variants.”
He further commented, “However with our new study approach, ee can now study what happens when many other proteins in the body are cleaved by Mpro during infection."
For the latest
SARS-CoV-2 Research, keep on logging to Thailand
Medical News.

