BREAKING! Study Reveals Compartmentalized SARS-CoV-2 Infection In The Human Body According To Tissue Types, Providing Evidence Of Viral Reservoirs!
COVID-19 News - Compartmentalized SARS-CoV-2 Infection In The Human Body Feb 03, 2023 3 years, 3 months, 2 weeks, 3 days, 3 hours, 51 minutes ago
COVID-19 News: A new study by researchers from Harvard University has revealed compartmentalized SARS-CoV-2 infection in the human body according to cell and tissue types and with the virus itself evolving and mutating to sustain its existence in these sub-micro environments! The study findings besides showing the existence of viral reservoirs and also validating viral persistence, also has numerous implications in terms of changing what we know about Long COVID issues and also the dynamics of variant emergence!
To date, SARS-CoV-2 distribution and circulation dynamics in the human body are not well understood due to challenges in assessing genomic data from tissue samples.
The study team developed experimental and computational workflows for high-depth viral sequencing and high-resolution genomic analyses from formalin-fixed, paraffin-embedded tissues and applied them to 120 specimens from six subjects with fatal COVID-19.
It was found that to varying degrees, viral RNA is present in extrapulmonary tissues from all subjects.
Interestingly, the majority of the 180 viral variants identified within subjects are unique to individual tissue samples.
The study team found more high-frequency (>10%) minor variants in subjects with a longer disease course, with one subject harboring ten such variants, exclusively in extrapulmonary tissues.
Surprisingly, it was found that one tissue-specific high-frequency variant even had a nonsynonymous mutation in the furin-cleavage site of the spike protein.
The study findings suggest adaptation and/or compartmentalized infection, illuminating the basis of extrapulmonary COVID-19 symptoms and potential for viral reservoirs, and have broad utility for investigating human pathogens.
The study findings were published in the peer reviewed journal: Nature Communications.
https://www.nature.com/articles/s41467-022-34256-y
In a
COVID-19 News coverage a year ago, we covered a German and British study that also showed that different variants of the SARS-CoV-2 virus hidden in different parts of the human body as the virus evolves to adapt for viral persistence in that particular part of the body and to evade the immune responses in that particular tissues or region.
https://www.thailandmedical.news/news/breaking-german-and-uk-research-indicates-that-people-infected-with-sars-cov-2-may-have-different-variants-hidden-in-various-parts-of-the-body
The study team utilized molecular tools to characterize the anatomical distribution of SARS-CoV-2 and investigate its evolution and circulation during natural human infection.
Using multiple RT-qPCR assays, high-depth sequencing, and fine resolution genomic analyses, the study team were able to shed light on the basis of tissue
-specific pathologies, observe trends that differentiated patients with shorter and longer disease courses, and identify extrapulmonary tissues that harbor compartmentalized infection later in disease.
The research was made possible by the study team’s unique development and validation of enhanced methods to sequence viral genomes from formalin-fixed, paraffin-embedded (FFPE) tissue specimens.
Viral sequencing and genomic analyses from tissues present additional challenges over fluid samples (e.g. blood, urine, swabs), particularly high host RNA background, but enrichment for viral sequence through hybrid capture enabled high-depth genome sequencing.
Even with the chemical crosslinking in FFPE samples, the study team found that viral genome coverage obtained from FFPE samples was comparable to that from frozen tissues.
The study findings also showed that minor variant identification sensitivity is consistently reliable above 500x mean depth of coverage, but sensitivity drops considerably at lower coverage depths.
Although this is notable for any study of intra-host variation, it is particularly vital to keep in mind for studies comparing variant profiles across samples with vastly different viral loads and corresponding coverage depths.
The study team demonstrated the utility and practical relevance of this method by deploying it to assemble viral genomes from dozens of tissue samples from six fatal cases of COVID-19, the majority of which had >500x coverage depth. There was very little SARS-CoV-2 variation in consensus-level genome assemblies between tissues in the same individual, underscoring the importance of accurate minor variant calling to characterize in vivo evolution.
The study findings identified viral genomic and transcriptomic commonalities and differences between subjects, and found indications of latency and compartmentalization that were particularly associated with longer infection times.
Though heterogeneity was observed in the tissues affected, each subject had relatively high viral loads in several extrapulmonary tissues, and viral loads were generally lower in subjects with a longer disease course.
The study findings showed that N gene RNA abundance, but not the proportion of N gene sgRNA reads, also decreased with a longer time interval post symptom onset.
This implies a down-regulation of viral structural components specifically later in the disease course, which could play a role in immune evasion.
Viral suppression of short sgRNA expression has been demonstrated in the context of SARS-CoV-1, as has regulation of sub-genomic RNA synthesis.
https://www.sciencedirect.com/science/article/pii/S1931312814003424
https://www.sciencedirect.com/science/article/pii/S002228360901167X
Such study findings raise the possibility that SARS-CoV-2 enters a slower replication state as infection continues.
The study team also identified more evidence of compartmentalization in subjects with longer times between symptom onset and death, with S03 and S04 subjects providing a striking comparison point. S03 had no tissue-specific high-frequency variants in any of the five extrapulmonary tissues profiled. This subject had only one day between symptom onset and death, and the short disease course, likely not providing sufficient time for compartmentalization to establish. By contrast, S04, with nine days between symptom onset and death, had 10 tissue-specific high-frequency variants, all in extrapulmonary tissues, strongly suggestive of compartmentalization.
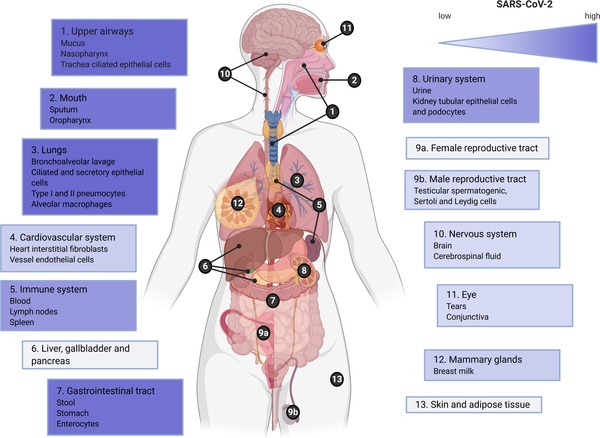
It should be noted that this evidence for compartmentalized infection, particularly in an immune-privileged site, has ramifications for SARS-CoV-2 evolution, as well as COVID-19 diagnostics and treatment.
Compartmentalized infection in the heart of S04 was consistent with the very high viral load by RT-qPCR and strong histopathological evidence for infection of the cardiomyocytes in this subject, as well as findings from previous reports of SARS-CoV-2 in the heart.
https://jamanetwork.com/journals/jamacardiology/article-abstract/2768914
These findings add to the credence of the hypothesis that direct infection may lead to the heart-related sequelae observed in some COVID-19 patients.
The testis, which also had evidence of compartmentalized infection in S04, is a known immune-privileged site and has been demonstrated to harbor compartmentalized infections of various pathogens, with implications for fertility and sexual transmission.
https://reproductive-health-journal.biomedcentral.com/articles/10.1186/s12978-021-01172-1
The SARS-CoV-2 virus may establish a viral reservoir in these anatomical sites, leading to ongoing tissue-specific pathology, and/or eventual reactivation.
Importantly, a persistent infection, particularly in an immunosuppressed patient, provides an environment for sustained viral evolution, which can result in the emergence of variants within the infected host and has been hypothesized as a potential source for novel variants of concern.
https://www.nejm.org/doi/full/10.1056/NEJMc2031364
https://www.medrxiv.org/content/10.1101/2021.06.03.21258228v1
It was noted that the Spike Q675H mutation, an adaptive mutation that has emerged independently in several SARS-CoV-2 lineages, arose in a compartmentalized infection in one individual (who was immunocompromised) after just 9 days of infection.
https://www.biorxiv.org/content/10.1101/2021.10.27.466055v1
https://www.mdpi.com/1999-4915/13/9/1801
Also, the emergence of a functionally advantageous mutation in the heart in a short period of time highlights the potential role of extra-pulmonary reservoirs of SARS-CoV-2 in viral evolution within persistently infected hosts.
The molecular methods deployed in this study for quantifying and sequencing viral RNAs can be used to characterize viral distribution and in vivo dynamics. For example, comparing variant profiles across viral populations sequenced from different compartments may reveal tissue-specific mutations that could be involved in invasion and replication in a specific tissue or cell type, or could reflect that that tissue harbors a compartmentalized infection.
Compartmentalized infection, particularly in an immune privileged site, may accommodate persistent infections, particularly in immunocompromised individuals, which fosters the generation of many novel mutations. Additionally, these sites could provide a viral reservoir that leads to reactivation or recrudescence. This phenomenon occurs across several viral families including HIV that establishes a reservoir in the central nervous system, flaviviruses remaining in the kidneys and testis, and filoviruses persisting in the testis and other compartments.
To date, observations of long-lasting symptoms and apparent recrudescence in COVID-1922 suggest the presence of extrapulmonary reservoirs for SARS-CoV-2, although the location of these reservoirs is unknown.
Although in vitro and animal models of infection can provide insights into many aspects of virology and host pathology, neither can fully recapitulate viral invasion and infection of multiple tissue compartments in systemic human disease.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7350325/
The study team hope that more studies will be focused on this area as it provides a better insight for developing treatment protocols for Long COVID and also ways to eradicate viral persistence.
For the latest
COVID-19 News, keep on logging in to Thailand Medical News.
Read Also:
https://www.thailandmedical.news/news/breaking-covid-19-news-finally,-u-s-nih-warns-sars-cov-2-viral-persistence-is-a-serious-issue-that-is-also-affecting-long-covid
https://www.thailandmedical.news/news/covid-news-new-york-scientists-find-sars-cov-2-viral-presence-in-the-lung-up-to-359-days-after-acute-phase-of-disease-even-in-the-negative-tested
https://www.thailandmedical.news/news/breaking-researchers-present-case-reports-of-long-covid-patients-with-persistence-of-residual-sars-cov-2-antigen-and-rna-in-various-tissues
https://www.thailandmedical.news/news/breaking-many-long-covid-19-patients-have-higher-levels-of-circulating-sars-cov-2-viral-rna-compared-to-those-with-acute-infection
https://www.thailandmedical.news/news/new-covid-19-diagnostics-urgently-needed-as-the-current-nasal-or-saliva-swabs-tests-are-unable-to-detect-viral-persistence-let-alone-new-variants
https://www.thailandmedical.news/news/breaking-latest-u-s-nih-study-reveals-viral-persistence-is-a-major-issue-and-that-sars-cov-2-replicates-in-tissues-for-months-in-post-covid-individua
https://www.thailandmedical.news/news/yet-another-study-confirms-viral-persistence-in-so-called-covid-19-recovered-patients-this-time-in-breast-and-appendix-tissues-of-two-patients-with-lo
https://www.thailandmedical.news/news/breaking-corpse-test-positive-for-sars-cov-2-virus-for-28-times-six-weeks-after-death-indicating-long-term-viral-persistence
https://www.thailandmedical.news/news/breaking-brazilian-and-american-study-discovers-that-the-testes-of-males-could-be-viral-sanctuaries-for-the-sars-cov-2-virus
https://www.thailandmedical.news/news/breaking-u-s-nih-study-shockingly-reveals-sars-cov-2-viral-persistence-throughout-human-body-and-in-the-brain-even-in-those-who-were-asymptomatic

