COVID-19 Research: University Of California Scientist Identify Rapidly Evolving Immune Evasion Protein SARS-CoV-2 ORF8
COVID-19 Research: Scientists from the Department of Molecular and Cell Biology, University of California-Berkeley have in a new study identified the SARS-CoV-2 ORF8 protein as being responsible for the rapidly evolving immune evasion characteristics of emerging strains and also responsible for the increasing pathogenicity of the coronavirus.
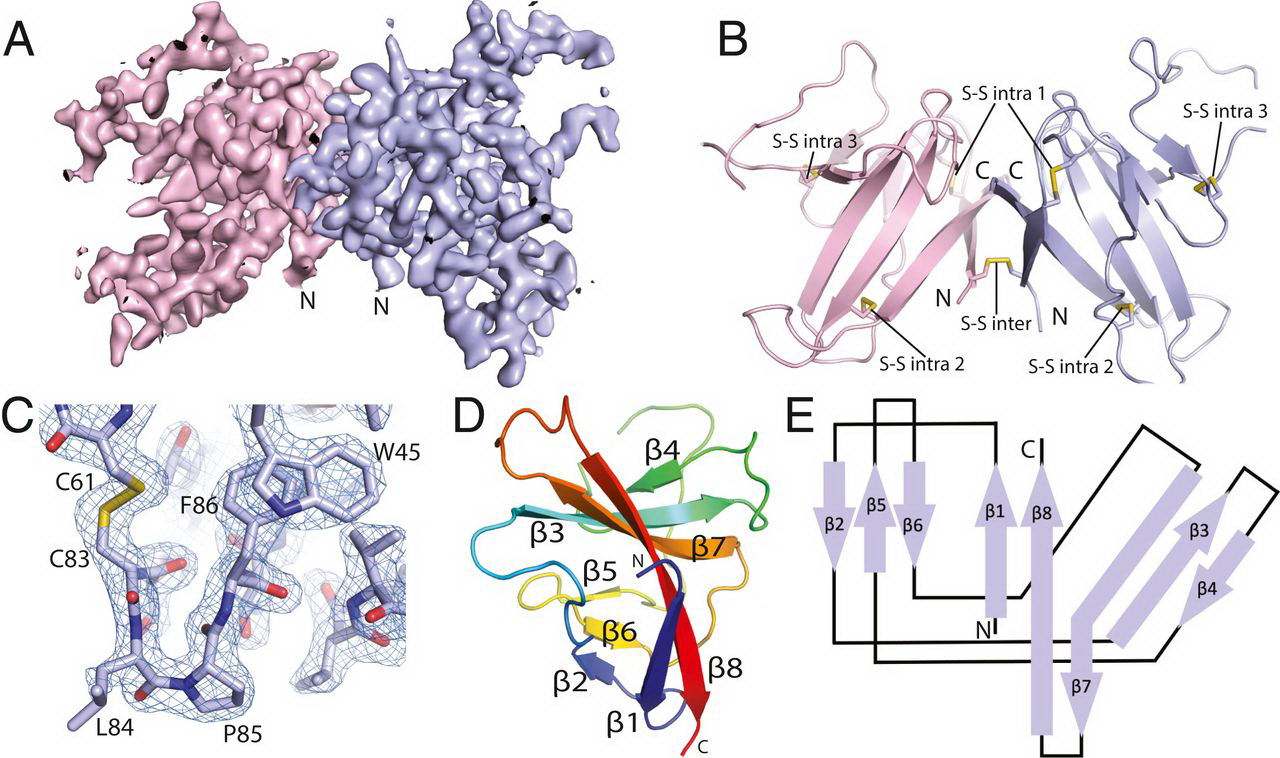 Crystal structure of SARS-CoV-2 ORF8. (A) A 2Fo-Fc electron density map of SARS-CoV-2 ORF8 crystallographic dimer determined to 2.04 Å (chain A, light blue; chain B, light pink). (B) Cartoon representation of the SARS-CoV-2 ORF8 crystallographic dimer. Disulfide bonds are modeled showing both intermolecular and intramolecular bond pairs. (C) Representative 2Fo-Fc density of the Cys83−Leu84−Pro85 turn motif. The map is contoured at 2σ and represented as a blue mesh. (D) Cartoon representation of the SARS-CoV-2 ORF8 monomer. β-strands are labeled β1 to β8, and chain is colored by rainbow gradient for clarity. (E) Topographic representation of the ORF8 monomer showing antiparallel β-sheets formed by β1 to β8.
Crystal structure of SARS-CoV-2 ORF8. (A) A 2Fo-Fc electron density map of SARS-CoV-2 ORF8 crystallographic dimer determined to 2.04 Å (chain A, light blue; chain B, light pink). (B) Cartoon representation of the SARS-CoV-2 ORF8 crystallographic dimer. Disulfide bonds are modeled showing both intermolecular and intramolecular bond pairs. (C) Representative 2Fo-Fc density of the Cys83−Leu84−Pro85 turn motif. The map is contoured at 2σ and represented as a blue mesh. (D) Cartoon representation of the SARS-CoV-2 ORF8 monomer. β-strands are labeled β1 to β8, and chain is colored by rainbow gradient for clarity. (E) Topographic representation of the ORF8 monomer showing antiparallel β-sheets formed by β1 to β8.
The structure of the SARS-CoV-2 ORF8 protein reveals two novel intermolecular interfaces layered onto an ORF7 fold. One is mediated by a disulfide bond, the other is noncovalent, and both are novel with respect to SARS-CoV. The structural analysis here establishes a molecular framework for understanding the rapid evolution of ORF8, its contributions to COVID-19 pathogenesis, and the potential for its neutralization by antibodies.
The team of HIV researchers, cellular biologists, and biophysicists who banded together to support COVID-19 science determined the atomic structure of a coronavirus protein thought to help the pathogen evade and dampen response from human immune cells.
Importantly the structural map which is now published in the journal PNAS, but has been open-access for the scientific community since August has laid the groundwork for new antiviral treatments tailored specifically to SARS-CoV-2, and enabled further investigations into how the newly emerged virus ravages the human body.
https://www.pnas.org/content/118/2/e2021785118
The molecular basis for the severity and rapid spread of the COVID-19 disease caused by SARS-CoV-2 is largely unknown. ORF8 is a rapidly evolving accessory protein that has been proposed to interfere with immune responses. The crystal structure of SARS-CoV-2 ORF8 was determined at 2.04-Å resolution by X-ray crystallography. The structure reveals a ∼60-residue core similar to SARS-CoV-2 ORF7a, with the addition of two dimerization interfaces unique to SARS-CoV-2 ORF8. A covalent disulfide-linked dimer is formed through an N-terminal sequence specific to SARS-CoV-2, while a separate noncovalent interface is formed by another SARS-CoV-2−specific sequence, 73YIDI76.
Together, the presence of these interfaces shows how SARS-CoV-2 ORF8 can form unique large-scale assemblies not possible for SARS-CoV, potentially mediating unique immune suppression and evasion activities.
Utilizing
X-ray crystallography, the study team built an atomic model of ORF8, and it highlighted two unique regions: one that is only present in SARS-CoV-2 and its immediate bat ancestor, and one that is absent from any other coronavirus.
Dr James Hurley, Lead Author, University of California-Berkeley Professor and Former Faculty Scientist, Lawrence Berkeley National Laboratory (Berkeley Lab) told Thailand Medical News, “These regions stabilize the protein which is a secreted protein, not bound to the membrane like the virus's characteristic spike proteins and create new intermolecular interfaces. The team and others in the research community, believe these interfaces are involved in reactions that somehow make SARS-CoV-2 more pathogenic than the strains it evolved from."
Typically generating protein structure maps is always labor intensive, as scientists have to engineer bacteria that can pump out large quantities of the molecule, manipulate the molecules into a pure crystalline form, and then take many, many X-ray diffraction images of the crystals. These images – produced as X-ray beams bounce off atoms in the crystals and pass through gaps in the lattice, generating a pattern of spots are combined and analyzed via special software to determine the location of every individual atom. This painstaking process can take years, depending on the complexity of the protein.
Also for many proteins, the process of building a map is helped along by comparing the unsolved molecule's structure to other proteins with similar amino acid sequences that have already been mapped, allowing scientists to make informed guesses about how the protein folds into its 3D shape.
However for ORF8, the study team had to start from scratch. ORF8's amino acid sequence is so unlike any other protein that scientists had no reference for its overall shape, and it is the 3D shape of a protein that determines its function.
Dr Hurley and his UC Berkeley colleagues, experienced in structural analysis of HIV proteins, worked with Dr Marc Allaire, a biophysicist and crystallography expert at the Berkeley Center for Structural Biology, located at Berkeley Lab's Advanced Light Source (ALS).
Together, the study team worked in overdrive for six months and Dr Hurley's lab generated crystal samples and passed them to Dr Allaire, who would use the ALS's X-ray beamlines to take the diffraction images. It took hundreds of crystals with multiple versions of the protein and thousands of diffraction images analyzed by special computer algorithms to puzzle together ORF8's structure.
Dr Hurley explained, "Coronaviruses mutate differently than viruses like influenza or HIV, which quickly accumulate many little changes through a process called hypermutation. In coronaviruses, big chunks of nucleic acids sometimes move around through recombination."
He added, “When this happens, big, new regions of proteins can appear. Genetic analyses conducted very early in the SARS-CoV-2 pandemic revealed that this new strain had evolved from a coronavirus that infects bats, and that a significant recombination mutation had occurred in the area of the genome that codes for a protein, called ORF7, found in many coronaviruses. The new form of ORF7, named ORF8, quickly gained the attention of virologists and epidemiologists because significant genetic divergence events like the one seen for ORF8 are often the cause of a new strain's virulence.”
Dr Hurley further added, "Basically, this mutation caused the protein to double in size, and the stuff that doubled was not related to any known fold. There's a core of about half of it that's related to a known fold type in a solved structure from earlier coronaviruses, but the other half was completely new."
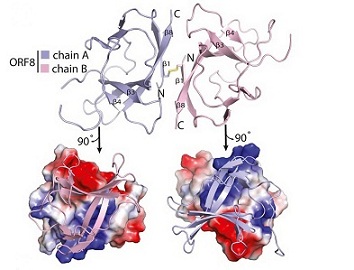
Similar to so many scientists working on COVID-19 research, Dr Hurley and his colleagues opted to share their findings before the data could be published in a peer-reviewed journal, allowing others to begin impactful follow-up studies months earlier than the traditional publication process would have allowed.
As Dr Allaire explained, the all-hands-on-deck crisis caused by the pandemic shifted everyone in the research community into a pragmatic mindset. Rather than worrying about who accomplished something first, or sticking to the confines of their specific areas of study, scientists shared data early and often, and took on new projects when they had the resources and expertise needed.
In this case, Dr Hurley's UC Berkeley co-authors had the viral protein and crystallography expertise, and Dr Allaire, a longtime collaborator, was right up the hill, also with crystallography expertise and, critically, a beamline that was still operational. The ALS had received special funding from the CARES Act to remain operational for COVID-19 investigations. The team knew from reviewing the SARS-CoV-2 genomic analysis posted in January that ORF8 was an important piece of the (then much hazier) pandemic puzzle, so they set to work.
The study team have since all moved on to other projects, satisfied that they laid the groundwork for other groups to study ORF8 in more detail. (Currently, there are several investigations underway focused on how ORF8 interacts with cell receptors and how it interacts with antibodies, as infected individuals appear to produce antibodies that bind to ORF8 in addition to antibodies specific to the virus's surface proteins.)
Dr Allaire, who is part of Berkeley Lab's Molecular Biophysics and Integrated Bioimaging Division added, "When we started this, other projects had been put on hold, and we had this unique opportunity to hunker down and solve an urgent problem. We worked very closely, with a lot of back and forth, until we got it right. It really has been one of the best collaborations of my career."
The study team concluded, “As structures are determined of ORF8 complexes with host receptors, we would expect to see ORF8 bound in these complexes as a covalent dimer. It is less certain whether to expect the YIDI motif-mediated noncovalent dimer interface to persist in some functional complexes. Given the high solvent exposure of these hydrophobic residues and their uniqueness to SARS-CoV-2 and its closest relatives, it is reasonable to expect them to be important for pathogenesis. It may be that these residues engage directly with host targets in some cases, which would presumably not be compatible with the persistence of the noncovalent ORF8 dimer. Alternatively, other regions may mediate direct interactions with receptors, while the YIDI interface promotes higher-order assembly and increases the avidity of binding to receptor oligomers, or induces the oligomerization of receptors. It is intriguing that ORF8 antibodies are major serological markers of SARS-CoV-2 infection. It will be important to define the immunogenic epitopes on SARS-CoV-2 ORF8, and whether they include the YIDI motif. To the extent that antibodies react with putative functional surfaces of ORF8 such as the YIDI motif, it will be important to determine whether the antibodies are neutralizing with respect to the pathogenic functions of ORF8 and, if so, whether these properties can be exploited therapeutically.”
For the latest on
COVID-19 Research, keep on logging to Thailand Medical News.

