Scientist From The University Of Wisconsin Discover That Genetic Variants Of Host ACE2 Are Differentially Identified By SARS-CoV-2
A new study by experts from the Department of Biochemistry of the University of Wisconsin-Madison reveals that the genetic variants of human angiotensin 2 or
ACE2 receptors can have altered recognition by SARS-CoV-2 coronavirus, the causative agent COVID-19) pandemic.
.jpg)
Comprehending how human ACE2 genetic variants differ in their recognition by SARS-CoV-2 can have a major impact in leveraging ACE2 as an axis for treating and preventing COVID-19.
In this study, the team interrogates thousands of ACE2 mutants to identify over one hundred human single-nucleotide variants (SNVs) that are likely to have altered recognition by the novel coronavirus, and make the complementary discovery that ACE2 residues distant from the spike interface can have a strong influence upon the ACE2-spike interaction. These study findings illuminate new links between ACE2 sequence and spike recognition, and will find wide-ranging utility in SARS-CoV-2 fundamental research, epidemiological analyses, and clinical trial design.
The study findings were published on a preprint server and have yet to be peer-reviewed.
https://www.biorxiv.org/content/10.1101/2020.09.17.301861v1
The human to human transmission and infectivity of SARS-CoV-2 primarily depends on the interaction between the receptor-binding domain (RBD) of viral spike protein and ACE2, a receptor expressed on the epithelial cells that line the human respiratory tract.
Hence, any studies deciphering the variability of spike-ACE2 interaction across the human population is of critical importance in terms of developing appropriate therapeutics and drugs or vaccines to contain the progression of the COVID-19 pandemic.
The study team from the University of Wisconsin-Madison tried to determine whether the SARS-CoV-2 spike protein identifies the single-nucleotide variants of the human ACE2 receptor differentially.
In order to address this query, the team conducted deep mutational scanning to evaluate how more than 3,500 amino acid substitutions in the extracellular peptidase domain of the ACE2 receptor may influence the interaction with the viral spike protein.
Based on the results by the scanning data, the study team found many novel ACE2 residues that impact the spike binding pattern and identified more than 100 single-nucleotide variants of ACE2 that are likely to be differentially recognized by the viral protein.
Significantly, the study team identified 597 locations in the ACE2’s extracellular domain containing 3,571 amino acid substitutions.
Interestingly of these substitutions, 68% were associated with reduced spike binding efficiency, and 4% were associated with increased spike binding efficiency. About 28% of the substitution had no statistically significant effect on the spike-ACE2 interaction.
Also, the team observed that these amino acid substitutions included 84% of all ACE2 missense single-nucleotide variants identified in the human population.
A grand total of 165 ACE2 variants were identified in the current study. The majority of these variants were distant from the spike-ACE2 interface, indic
ating that the spike-ACE2 interaction can be modulated by ACE2 mutations that are distal to the spike binding interface.
In order to validate the accuracy of current study data, the study team evaluated the binding impact of previously identified important ACE2 residues (S19, Q24, D30, H34, D38, Y41, Q42, Y83, and K353) that are present at the interface between the viral spike protein and the α-helical and β-turn sequences of ACE2. In agreement with previous study findings, they observed that these residues play essential roles in mediating the spike-ACE2 interaction.
An important aspect of the study is the identification of previously unrecognized ACE2 residues that are crucial for spike binding. Precisely, the scientists observed that some residues adjacent to the chloride-binding domain of ACE2 play vital roles in modulating the spike-ACE2 interaction by altering the structural conformation of ACE2.
The study team by analyzing the allele frequency of each ACE2 variant in the general population, estimated that about 320 to 365 of 100,000 individuals carry ACE2 variants that may reduce the spike binding; whereas, 4 to 12 of 100,000 individuals carry ACE2 variants that may increase the spike binding.
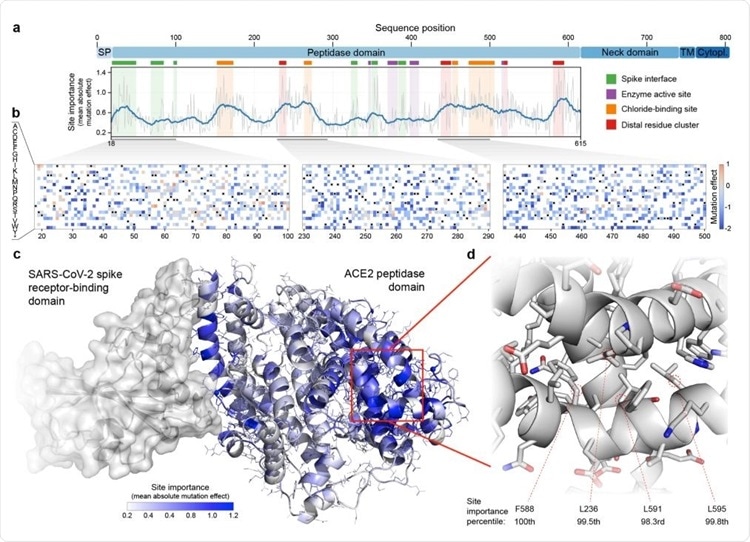 Large-scale mutagenesis of ACE2’s peptidase domain. (a) Analysis of how amino acid substitutions across positions 18-615 affect binding of the SARS-CoV-2 spike protein. The plot quantifies the importance of each site by taking the mean of the absolute value of all mutation effects observed at that site. The grey line represents the mean absolute value of the mutation effect and the blue line shows the moving average to highlight general regions of ACE2 that are important for binding. Key structural landmarks are highlighted with shaded regions along the length of the sequence. (b) Mutation effect heat maps for three different regions of ACE2. Red denotes mutations that increase ACE2 spike binding; blue denotes reduced binding. Overall, we observe the effects of 3571 amino acid substitutions across 597 positions in ACE2’s peptidase domain. (c) The mean absolute mutation effect mapped onto the threedimensional ACE2 structure (PDB ID: 6LZG). Residues near the spike interface are important for binding, in addition to many sites located on the distal lobe of the protein domain. (d) The most important region of ACE2 structure is composed of a tightly packed cluster of residues located over 30 Å from the spike interface.
Large-scale mutagenesis of ACE2’s peptidase domain. (a) Analysis of how amino acid substitutions across positions 18-615 affect binding of the SARS-CoV-2 spike protein. The plot quantifies the importance of each site by taking the mean of the absolute value of all mutation effects observed at that site. The grey line represents the mean absolute value of the mutation effect and the blue line shows the moving average to highlight general regions of ACE2 that are important for binding. Key structural landmarks are highlighted with shaded regions along the length of the sequence. (b) Mutation effect heat maps for three different regions of ACE2. Red denotes mutations that increase ACE2 spike binding; blue denotes reduced binding. Overall, we observe the effects of 3571 amino acid substitutions across 597 positions in ACE2’s peptidase domain. (c) The mean absolute mutation effect mapped onto the threedimensional ACE2 structure (PDB ID: 6LZG). Residues near the spike interface are important for binding, in addition to many sites located on the distal lobe of the protein domain. (d) The most important region of ACE2 structure is composed of a tightly packed cluster of residues located over 30 Å from the spike interface.
Of significance, the study team observed that the frequency of having specific variants that alter the spike binding varies between individuals with a different ancestral origin.
For instance, compared to the general population, the African population is five times more likely to carry specific ACE2 variants, such as p.Met82Ile.
With regards to the mode of action of ACE2 variants, the study revealed that the variants with reduced binding efficiency have a three-fold reduction in maximum binding signals than that of wild-type ACE2.
The ACE2 variants identified in the study were significantly different from wild-type ACE2 in terms of spike binding affinity, the number of receptors present on the cell membrane, and receptor turnover rate at the cell membrane.
The team commented, “The ability of SARS-CoV-2 virions to enter and infect human host cells is dependent upon a number of ACE2 properties: the binding affinity for the SARS-CoV-2 spike, the amount of ACE2 that migrates through the endoplasmic reticulum to reach the cell surface, and the turnover rate of ACE2 at the cell membrane. Our study findings has identified ACE2 SNVs that are markedly distinct from the wild-type protein with respect to the above properties, and thus present new opportunities to interleave ACE2 biochemical studies, ACE2 variant animal experiments, and observations of the human population. This work has also revealed that mutations in ACE2 residues distal to the SARS-CoV-2 spike-binding interface may alter ACE2 properties relevant to SARS-CoV-2 recognition; this finding will motivate future empirical and in silico research efforts to expand their scope beyond the spike binding interface. Collectively our results provide key insights that can be readily actioned toward fundamental research, clinical studies, and epidemiological analyses for the treatment and prevention of SARS-CoV-2 infection.”
The research findings also pave the way toward the identification of novel therapeutic targets and prospective drugs.
.jpg)
